MP2 DNA Base Stacking Energy: A Computational Guide for Structural Biology & Drug Discovery
This comprehensive guide details the use of second-order Møller-Plesset perturbation theory (MP2) for calculating DNA base pair stacking interactions.
MP2 DNA Base Stacking Energy: A Computational Guide for Structural Biology & Drug Discovery
Abstract
This comprehensive guide details the use of second-order Møller-Plesset perturbation theory (MP2) for calculating DNA base pair stacking interactions. Tailored for computational chemists, biophysicists, and pharmaceutical researchers, we explore the fundamental role of π-π stacking in DNA stability, provide practical methodologies for MP2 calculations, address common computational challenges, and validate MP2 against advanced methods like CCSD(T) and DFT-D. The article equips researchers with the knowledge to accurately model these critical non-covalent forces, directly informing rational drug design and the understanding of genetic diseases.
DNA Stacking 101: Why π-π Interactions and MP2 Theory Are Crucial for Stability
The Central Role of Base Stacking in DNA Structure and Energetics
Base stacking, the attractive, non-covalent interaction between adjacent nucleobases in DNA, is a fundamental determinant of duplex stability, rigidity, and biological function. Within the broader thesis on MP2 Calculation for DNA Base Pair Stacking Research, this application note establishes the experimental and computational context. High-level ab initio methods like MP2 (Møller-Plesset perturbation theory to second order) are crucial for accurately calculating stacking interaction energies, as they account for electron correlation effects essential for describing dispersion forces—the dominant component of stacking energetics. These quantum mechanical (QM) benchmarks inform and validate force fields used in molecular dynamics (MD) simulations of drug-DNA interactions.
Table 1: Calculated Stacking Interaction Energies for Common Dinucleotide Steps (in vacuo, MP2/cc-pVDZ level approximations)
| Dinucleotide Step | Stacking Energy (ΔE, kcal/mol) | Dominant Contribution | Notes |
|---|---|---|---|
| 5'-CpG-3' / 5'-CpG-3' (CG) | -14.2 to -16.5 | Dispersion | Most stable step; significant sequence-dependent variability. |
| 5'-GpC-3' / 5'-GpC-3' (GC) | -12.8 to -14.1 | Dispersion | |
| 5'-ApA-3' / 5'-TpT-3' (AA/TT) | -9.5 to -11.2 | Dispersion + Electrostatics | Roll and twist deform easily. |
| 5'-ApT-3' / 5'-ApT-3' (AT) | -8.0 to -9.5 | Electrostatics | Weaker stacking, easier unstacking. |
| 5'-TpA-3' / 5'-TpA-3' (TA) | -6.5 to -8.5 | Electrostatics | Least stable step; a potential kink site. |
Table 2: Comparison of Computational Methods for Stacking Energy Calculation
| Method | Speed | Accuracy for Stacking | Key Limitation for DNA |
|---|---|---|---|
| MP2 | Moderate | High | Basis set superposition error (BSSE); cost scales as O(N⁵). |
| DFT-D3 (w/ dispersion correction) | Fast | Moderate to High | Dependent on functional choice; may misbalance interactions. |
| CCSD(T) (Gold Standard) | Very Slow | Very High | Prohibitively expensive for large systems. |
| Molecular Mechanics (MM) | Very Fast | Low (without QM parametrization) | Force field dependent; poor transferability. |
Experimental Protocols for Correlative Studies
Protocol 3.1: Measuring Stacking Thermodynamics via Differential Scanning Calorimetry (DSC)
Objective: To determine experimentally the enthalpy (ΔH) and melting temperature (Tm) of DNA duplex formation, which correlates with overall stability influenced by base stacking. Materials: See Scientist's Toolkit. Procedure:
- Sample Preparation: Prepare high-purity DNA oligonucleotides in matched buffer (e.g., 10 mM sodium phosphate, 100 mM NaCl, pH 7.0). Degas solution to prevent bubble artifacts.
- Instrument Setup: Equilibrate DSC cell at 15°C. Load sample and reference (buffer) cells. Set scan rate to 1°C/min from 15°C to 95°C.
- Data Acquisition: Perform a buffer-buffer baseline scan. Replace reference with buffer, load sample, and run the temperature scan.
- Data Analysis: Subtract baseline from sample scan. Integrate the area under the heat capacity peak to obtain ΔH. Identify Tm at the peak maximum. Deconvolute data for nearest-neighbor parameters using specialized software.
Protocol 3.2: Probing Stacking-Dependent Conformation via Circular Dichroism (CD) Spectroscopy
Objective: To characterize the secondary structure (B-DNA, A-DNA, Z-DNA) and stacking organization of a DNA duplex. Procedure:
- Sample Prep: Dilute DNA to an OD260 of ~0.5-1.0 in appropriate buffer.
- Instrument Calibration: Calibrate CD spectropolarimeter with ammonium d-camphor-10-sulfonate.
- Spectrum Acquisition: Use a 1 cm pathlength quartz cuvette. Scan from 320 nm to 200 nm at 20 nm/min, 1 nm bandwidth, 1 s response time. Average 3 scans.
- Analysis: B-DNA shows a positive band at ~275 nm and negative band at ~245 nm. Increased positive band intensity often indicates stronger base stacking. Plot mean residue ellipticity [θ] vs. wavelength.
Protocol 3.3: Computational Workflow for MP2 Benchmarking of Stacking Energies
Objective: To calculate ab initio stacking interaction energies for a dinucleotide step. Procedure:
- Geometry Extraction: Extract coordinates for a dinucleotide step (e.g., CpG) from a high-resolution crystal structure (PDB ID: 1BNA).
- Model Preparation: Remove backbone, cap nucleobases with methyl groups at sugar attachment points. Ensure no close van der Waals contacts.
- Geometry Optimization: Optimize the geometry of the stacked dimer and individual monomers using DFT (e.g., B3LYP/6-31G(d)) to create a realistic starting point.
- Single-Point MP2 Calculation: Perform a high-level single-point energy calculation on the optimized geometry using MP2 with a medium-sized basis set (e.g., cc-pVDZ).
- BSSE Correction: Apply the Counterpoise Correction method to correct for Basis Set Superposition Error.
- Energy Calculation: Compute the stacking interaction energy: ΔEstack = Edimer - (EmonomerA + EmonomerB).
Visualization of Concepts and Workflows
Diagram Title: MP2 to Drug Design Pipeline
Diagram Title: DSC Experimental Workflow
The Scientist's Toolkit
Table 3: Key Research Reagent Solutions & Materials
| Item | Function / Application | Notes |
|---|---|---|
| High-Purity DNA Oligonucleotides | Substrate for all biophysical and computational studies. | HPLC or PAGE purified; essential for accurate thermodynamics. |
| Sodium Phosphate Buffer (with NaCl) | Standard buffer for DNA studies; controls ionic strength. | 10 mM phosphate, 100 mM NaCl, pH 7.0 mimics physiological conditions. |
| Differential Scanning Calorimeter (DSC) | Measures heat changes during DNA melting; provides direct ΔH. | Requires degassed, matched samples. |
| Circular Dichroism Spectropolarimeter | Probes chiral environment of nucleobases; reports on stacking geometry. | Uses quartz cuvettes; sensitive to buffer absorbance. |
| Quantum Chemistry Software (e.g., Gaussian, GAMESS, ORCA) | Performs MP2 and other ab initio calculations. | Requires high-performance computing (HPC) resources. |
| Molecular Dynamics Software (e.g., AMBER, GROMACS, NAMD) | Simulates DNA dynamics and drug binding using force fields. | Force fields (e.g., parmBSC1) are parameterized using QM data. |
| BSSE-Corrected Basis Sets (e.g., cc-pVDZ, aug-cc-pVDZ) | Basis functions for QM calculations; balance of accuracy and cost. | Correlation-consistent polarized basis sets are recommended. |
Application Notes
The quantification of hydrogen bonding in DNA base pairs is a standard biophysical metric. However, a comprehensive understanding of pairing fidelity and stability, especially within the context of stacking interactions in duplex DNA, requires decomposition of the total interaction energy into its fundamental physical components. This is critical for research in mutagenesis, drug design targeting specific DNA sequences, and the engineering of nucleic acid nanostructures. Within a broader thesis employing MP2-level ab initio calculations for DNA base pair stacking research, this protocol details the application of Energy Decomposition Analysis (EDA) schemes to isolate and quantify the contributions of Pauli repulsion, electrostatic interaction, dispersion, and orbital mixing (charge transfer, polarization) to base pair binding.
Key Quantitative Data from Representative MP2/EDA Studies Table 1: Energy Decomposition Analysis (kcal/mol) for a Canonical Guanine-Cytosine (GC) Base Pair (Watson-Crick) using a DFT-based EDA scheme (PBE0-D3/def2-TZVP) as a reference for MP2 trends. Geometry optimized at MP2/cc-pVDZ.
| Energy Component | Contribution (kcal/mol) | Physical Interpretation |
|---|---|---|
| Electrostatics (ΔE_ele) | -81.2 | Attraction due to permanent charge distributions (e.g., H-bond dipoles). |
| Pauli Repulsion (ΔE_Pauli) | +102.5 | Repulsion between occupied orbitals enforcing molecular shape. |
| Dispersion (ΔE_disp) | -25.8 | Attraction from correlated electron fluctuations (critical for stacking). |
| Orbital Interaction (ΔE_oi) | -45.3 | Stabilization from charge transfer & polarization (e.g., H-bond covalency). |
| Total Interaction (ΔE_int) | -49.8 | Sum of all components (ΔEele + ΔEPauli + ΔEdisp + ΔEoi). |
Table 2: Comparison of EDA Components for GC vs. AT Base Pairs (Model System: MP2/6-311G(0.25, 0.15) with SAPT reference).
| Base Pair | ΔE_ele | ΔE_Pauli | ΔE_disp | ΔE_oi | ΔE_int |
|---|---|---|---|---|---|
| G-C (WC) | -75.6 | +94.1 | -22.4 | -42.5 | -46.4 |
| A-T (WC) | -41.3 | +55.7 | -15.2 | -18.9 | -19.7 |
Experimental Protocols
Protocol 1: Geometry Optimization of Isolated Base Pairs for Subsequent EDA
- System Setup: Using computational chemistry software (e.g., Gaussian, ORCA, PSI4), define the coordinates for the two nucleobases (e.g., Guanine and Cytosine) in their Watson-Crick hydrogen-bonding alignment. Employ a standard dimer-centered coordinate system.
- Method Selection: Set the calculation method to MP2. Select a correlation-consistent basis set (e.g., cc-pVDZ) or a Pople-type basis set with diffuse and polarization functions (e.g., 6-311++G(d,p)).
- Optimization Run: Execute a geometry optimization calculation with tight convergence criteria on the forces and energy. Ensure no symmetry constraints are applied.
- Validation: Confirm the optimized structure possesses expected hydrogen bond lengths (N-H···N ~1.9 Å, N-H···O ~1.8 Å) and planar geometry. Perform a frequency calculation to verify the structure is a true minimum (no imaginary frequencies).
Protocol 2: Symmetry-Adapted Perturbation Theory (SAPT) Energy Decomposition at MP2 Reference Note: SAPT provides a physically rigorous decomposition directly, often used as a benchmark for DFT-based EDA.
- Input Preparation: Use the MP2-optimized geometry from Protocol 1. In the input file (e.g., for PSI4), specify the monomers by defining the atom indices belonging to base A and base B.
- Calculation Specification: Set the calculation to
SAPT(MP2)to use MP2 amplitudes for the intramonomer correlation. Specify a suitable basis set (e.g., aug-cc-pVDZ). TheSAPT_BASISkeyword can be set for more efficient calculations. - Decomposition Output: Run the calculation. The output will decompose the total interaction energy into explicit components:
- Electrostatics (Eelst): First-order electrostatic energy.
- Exchange (Eexch): Pauli repulsion (first-order exchange).
- Induction (Eind): Energy from polarization (can include charge transfer).
- Dispersion (Edisp): Second-order dispersion energy.
- δ(HF): A higher-order correction term.
- Analysis: Sum components:
E_int = E_elst + E_exch + E_ind + E_disp + δ(HF). Compare the magnitude of dispersion (E_disp) to the classical electrostatic term to assess the relative importance of non-electrostatic forces.
Protocol 3: DFT-Based Energy Decomposition Analysis (EDA) using the Amsterdam Modeling Suite (AMS) Note: This protocol uses the Morokuma-Ziegler-type EDA as implemented in the ADF module.
- Fragment Definition: Load the optimized dimer structure. Define the two nucleobases as separate, non-interacting fragments.
- Calculation Setup: Select the ADF module. Choose a hybrid functional (e.g., PBE0) with an empirical dispersion correction (e.g., Grimme's D3-BJ). Specify a high-quality triple-zeta basis set with polarization functions (e.g., TZ2P).
- EDA Request: In the "Properties" or "Detailed" settings, enable the "Energy Decomposition" (EDA) option.
- Execution & Interpretation: Run the calculation. The EDA results will decompose the instantaneous interaction energy (ΔEint) into:
- ΔEPauli: Pauli repulsion.
- ΔEelstat: Quasi-classical electrostatic interaction between fragments.
- ΔEoi: Orbital interaction (combining polarization and charge transfer).
- ΔEdisp: Dispersion energy (from the empirical correction). The sum equals ΔEint.
Mandatory Visualization
Title: Computational Workflow for Base Pair Energy Decomposition
Title: Components of Interaction Energy in EDA/SAPT
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Computational Tools for MP2-based Base Pair EDA
| Item / Software | Function / Description | Example / Specification |
|---|---|---|
| Quantum Chemistry Package | Primary engine for MP2 and EDA calculations. | Gaussian 16, ORCA 5.0, PSI4, Amsterdam Modeling Suite (AMS/ADF). |
| Basis Set Library | Mathematical functions describing electron orbitals; accuracy is critical. | cc-pVDZ, aug-cc-pVTZ (for SAPT), 6-311++G(d,p), TZ2P (in ADF). |
| Wavefunction Analysis Tool | Visualizes orbitals and electron density for interpreting ΔE_oi. | Multiwfn, VMD with plugins, Chemcraft. |
| Geometry Visualizer | Prepares inputs and analyzes optimized molecular structures. | Avogadro, GaussView, PyMOL. |
| High-Performance Computing (HPC) Cluster | Provides necessary CPU/core hours and memory for MP2 calculations. | Linux cluster with MPI/OpenMP support, ~64-256 GB RAM recommended for aug-cc-pVTZ. |
In computational studies of non-covalent interactions, such as the stacking of DNA nucleobases, accurately capturing dispersion forces is paramount. These interactions are crucial for maintaining the double-helix structure, influencing DNA-protein binding, and affecting the mechanisms of drug intercalation. While Density Functional Theory (DFT) with empirical dispersion corrections is popular, wavefunction-based methods provide a more rigorous, parameter-free framework. Among these, second-order Møller-Plesset perturbation theory (MP2) has long been considered the "gold standard" for its balanced description of dispersion at a computationally tractable cost for moderate-sized systems like base pairs. This note details its application within a research thesis focused on quantifying stacking interactions in DNA.
The Theoretical Hierarchy and MP2's Position
The accuracy of electronic structure methods for dispersion can be ranked by their theoretical rigor and computational cost. MP2 occupies a critical niche.
Table 1: Method Comparison for Dispersion in Base Pair Stacking
| Method | Description | Treatment of Dispersion | Approx. Cost (Scaling) | Typical Use Case in Stacking Studies |
|---|---|---|---|---|
| HF | Hartree-Fock | None (Fails to capture dispersion). | N⁴ | Not used for final dispersion energy. |
| DFT (GGA) | Generalized Gradient Approximation | Poor, often severely underestimated. | N³ | Not recommended for stacking. |
| DFT-D | DFT with empirical dispersion correction | Good, but relies on system-specific parameterization. | N³ to N⁴ | Screening studies on large fragments. |
| MP2 | Second-order Møller-Plesset Perturbation Theory | Excellent, from first principles. Captures a large fraction of dispersion. | N⁵ | Gold standard for benchmarking systems of 50-100 atoms. |
| CCSD(T) | Coupled-Cluster with Singles, Doubles, & perturbative Triples | Near-exact ("gold standard" for total energy). | N⁷ | High-level reference for small model systems (<30 atoms). |
| DLPNO-CCSD(T) | Domain-based Local Pair Natural Orbital CCSD(T) | Near-exact but with reduced cost via localization. | ~N⁴ to N⁵ | High-accuracy validation for MP2 on larger fragments. |
Core Protocol: Performing an MP2 Stacking Energy Calculation
This protocol outlines the steps to compute the stacking interaction energy between two DNA nucleobases (e.g., Adenine and Thymine) using MP2.
Objective: Calculate the CCSD(T)-level interaction energy at the complete basis set (CBS) limit using an MP2-based focal-point approach. Workflow Summary: A medium-sized basis set is used for the expensive CCSD(T) calculation, while the larger basis set effect and core correlation are captured via more affordable MP2 and MP2-F12 calculations.
Diagram Title: MP2-Based Focal-Point Approach for Stacking Energy
Detailed Protocol Steps:
- System Preparation & Geometry Optimization:
- Obtain initial coordinates for a stacked DNA base pair dimer (e.g., from a crystal structure, PDB ID: 1BNA).
- Software: Use
PSI4,Gaussian 16, orORCA. - Method: Optimize the geometry of the dimer and the isolated monomers using a cost-effective method like
ωB97X-D/6-31G*. This functional includes dispersion and is reliable for initial optimization. - Critical Check: Perform a frequency calculation (
freq=) on the optimized structures to confirm they are true minima (no imaginary frequencies).
Single-Point Energy Calculations (The Core):
- Using the optimized geometries, perform a series of single-point energy calculations at higher levels of theory.
- MP2 Series: Calculate energies for the dimer and monomers using MP2 with the
aug-cc-pVTZandaug-cc-pVQZbasis sets. These are diffuse-augmented correlation-consistent basis sets essential for describing weak intermolecular interactions. - High-Level Reference: Perform a CCSD(T) calculation with the
aug-cc-pVDZbasis set. Note: This is computationally demanding and may be restricted to smaller model systems or performed using the DLPNO approximation.
Basis Set Extrapolation (MP2/CBS):
- Use the MP2/aVTZ and MP2/aVQZ energies to extrapolate to the complete basis set (CBS) limit. A common two-point formula is:
E_MP2/CBS = (E_MP2/QZ * X_QZ^3 - E_MP2/TZ * X_TZ^3) / (X_QZ^3 - X_TZ^3)whereX_nis the cardinal number (TZ=3, QZ=4).
- Use the MP2/aVTZ and MP2/aVQZ energies to extrapolate to the complete basis set (CBS) limit. A common two-point formula is:
Interaction Energy Calculation & Focal-Point Analysis:
- For each level of theory, compute the stacking interaction energy:
ΔE = E_dimer - (E_monomer_A + E_monomer_B). - Apply the Counterpoise (CP) correction to each
ΔEto account for Basis Set Superposition Error (BSSE). This involves computing monomer energies in the full dimer basis set. - Final Focal-Point Energy: Combine the results to approximate a CCSD(T)/CBS energy:
ΔE_CCSD(T)/CBS ≈ ΔE_CCSD(T)/aVDZ + [ΔE_MP2/CBS - ΔE_MP2/aVDZ]This uses MP2 to approximate the effect of increasing the basis set for CCSD(T).
- For each level of theory, compute the stacking interaction energy:
Table 2: Key Computational "Reagents" for MP2 Stacking Studies
| Item/Software | Function & Relevance | Key Notes for Protocol |
|---|---|---|
| Quantum Chemistry Packages | ||
PSI4 |
Open-source suite. Excellent for automated CBS extrapolations and focal-point analyses. | Use psi4.energy("MP2/aug-cc-pVTZ"). Efficient for protocol automation. |
Gaussian 16 |
Industry standard. Robust and user-friendly with extensive model chemistry options. | Use # MP2/aug-cc-pVTZ Counterpoise=2. Well-documented for BSSE. |
ORCA |
Efficient, MPI-parallelized. Excellent for local correlation methods like DLPNO-CCSD(T). | Use ! DLPNO-CCSD(T) aug-cc-pVDZ aug-cc-pVDZ/C. For high-level validation. |
| Basis Sets | ||
aug-cc-pVnZ (aVnZ) |
The de facto standard for non-covalent interactions. "aug-" adds diffuse functions for dispersion. | n=D,T,Q. Larger n increases accuracy and cost. Essential for CBS extrapolation. |
jun-cc-pVnZ |
A newer family with improved cost-accuracy trade-off for MP2. | Can be more efficient than aug- sets for similar accuracy in stacking. |
| Model Geometries | ||
Protein Data Bank (PDB) |
Source of experimental DNA structures to extract initial base-pair stacking geometries. | Use structures like 1BNA. Extract coordinates for your specific dimer of interest. |
| Analysis Tools | ||
SAPT |
Symmetry-Adapted Perturbation Theory. Decomposes interaction energy (electrostatics, induction, dispersion). | Can be performed at the MP2 level (SAPT2) to quantify the exact dispersion contribution. |
ChemCraft / VMD |
Visualization software to analyze geometries, intermolecular distances, and electron density plots. | Critical for interpreting results and creating publication-quality figures. |
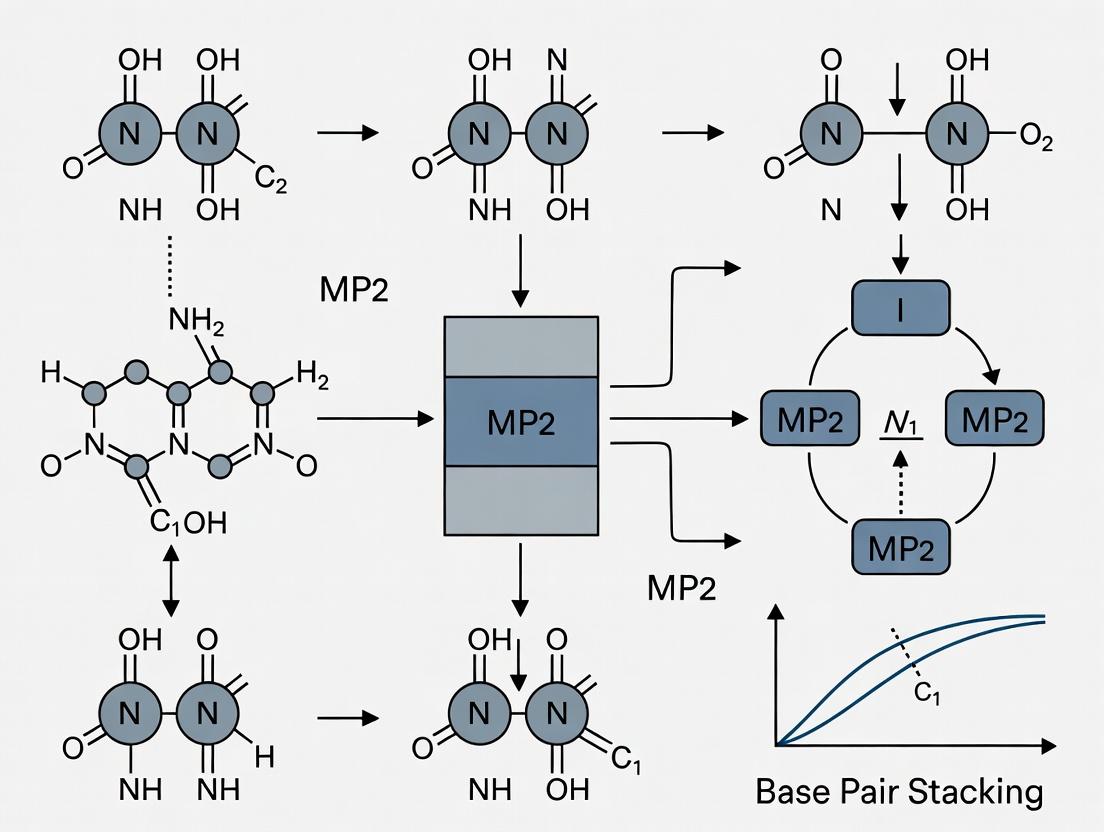
Visualizing the Role of MP2 in the Research Workflow
Diagram Title: MP2's Role in DNA Stacking Research Workflow
Within the broader thesis investigating DNA base pair stacking interactions using Møller-Plesset second-order perturbation theory (MP2), defining the precise geometric parameters of dinucleotide steps is fundamental. MP2 calculations, which account for electron correlation effects critical for describing dispersion forces in π-π stacking, require accurate starting geometries and parameters for meaningful energy comparisons. This application note details the definition, measurement, and application of the key dimeric stacking parameters: Step Parameters (Shift, Slide, Rise), Twist, and Roll/Tilt. These parameters are the principal descriptors of the relative orientation and displacement of two adjacent base pairs in a nucleic acid duplex, forming the quantitative basis for correlating geometry with stacking energy computed via high-level quantum mechanics.
Definitions of Key Geometrical Parameters
The geometry of a base pair step is described by six parameters: three rotational (Twist, Roll, Tilt) and three translational (Shift, Slide, Rise). They are defined by transforming one base pair (Bp1) onto the next (Bp2) via a coordinate frame centered on each base pair's midpoint and long axis.
- Twist: The rotation around the helical axis (z-axis of the step). It is the primary descriptor of helicity.
- Roll: The rotation around the long axis of the base pairs (x-axis), describing the "opening" angle between base pairs towards the major or minor groove.
- Tilt: The rotation around the short axis (y-axis), describing a "lifting" at one side of the base pair step.
- Shift: The translation along the short axis (y-axis, minor groove direction).
- Slide: The translation along the long axis (x-axis, direction from one backbone to the other). A key discriminator between A- and B-DNA forms.
- Rise: The translation along the helical axis (z-axis), representing the distance between base pairs.
The following table summarizes typical values for key step parameters in standard DNA conformations, as derived from crystallographic databases and used as benchmarks in MP2 stacking energy studies.
Table 1: Characteristic Step Parameters for Canonical DNA Forms
| Parameter | B-DNA (Canonical) | A-DNA | Protein-Bound B-DNA | Notes for MP2 Studies |
|---|---|---|---|---|
| Twist (°) | 36.0 ± 5.0 | 32.7 ± 4.0 | Variable, often >36° | High Twist can reduce orbital overlap, affecting dispersion energy. |
| Rise (Å) | 3.38 ± 0.2 | 2.56 ± 0.2 | ~3.3 Å | Rise is inversely correlated with stacking energy magnitude; critical for MP2 potential energy scans. |
| Slide (Å) | 0.0 ± 0.5 | -1.5 ± 0.4 | ~0.0 | Negative Slide is hallmark of A-form; directly influences base overlap and electrostatic components. |
| Roll (°) | 0.0 ± 5.0 | 5.0 ± 3.0 | Often positive (>+5°) | Positive Roll opens major groove; MP2 can quantify energy cost of deformation. |
| Tilt (°) | 0.0 ± 2.0 | 0.0 ± 2.0 | Variable | Usually small; significant tilt can indicate distortion. |
| Shift (Å) | 0.0 ± 0.5 | 0.0 ± 0.3 | Variable | Lateral displacement; can be sequence-dependent. |
Table 2: Example MP2/cc-pVDZ Stacking Energies vs. Slide Parameter for a G:C Step
| Slide (Å) | Twist (°) | Rise (Å) | MP2 Stacking Energy (kcal/mol) | Relative Stability |
|---|---|---|---|---|
| -2.0 | 36 | 3.4 | -12.1 | Low (A-like, unfavorable for B-DNA) |
| -1.0 | 36 | 3.4 | -15.4 | Intermediate |
| 0.0 | 36 | 3.4 | -17.8 | Most Stable (Canonical B) |
| +1.0 | 36 | 3.4 | -16.2 | Intermediate |
| +2.0 | 36 | 3.4 | -14.0 | Low (Over-stretched) |
Experimental Protocols
Protocol 1: Extracting Step Parameters from a PDB File
This protocol details the computational procedure to calculate dimeric stacking parameters from an atomic coordinate file (e.g., from X-ray crystallography or MD simulation snapshots), generating the necessary input for MP2 geometry optimization or single-point energy calculations.
Materials:
- Source PDB file of a nucleic acid structure.
- Software: 3DNA/Curves+ or MDANSE.
- Workstation with command-line interface.
Procedure:
- Structure Preparation: Isolate the DNA duplex of interest. Remove water, ions, and protein if present. Ensure base pairs are correctly paired. Hydrogen atoms may need to be added using molecular modeling software (e.g., UCSF Chimera).
- Software Execution (3DNA Example):
- Run the
find_paircommand to identify base pairs and their interacting steps: - Execute the
analyzecommand to calculate all step and helical parameters: - This generates
output.outandoutput.parms. The.parmsfile contains a table with all six step parameters (Shift, Slide, Rise, Tilt, Roll, Twist) for each dinucleotide step.
- Run the
- Data Curation: Extract the relevant step parameters. Average values over symmetrical steps or multiple copies in the asymmetric unit. Associate parameters with sequence context (e.g., AA/TT vs. CG/GC steps).
Protocol 2: Constructing a Geometry Scan for MP2 Stacking Energy Calculation
This protocol outlines the generation of a series of dimer geometries with systematic variation of a target parameter (e.g., Slide) for subsequent MP2 energy calculation, enabling the construction of energy profiles.
Materials:
- High-quality starting B-DNA dimer coordinates (e.g., from canonical fiber model).
- Quantum Chemistry Software: Gaussian, GAMESS, ORCA, or PySCF.
- Scripting environment (Python, Bash) for batch job generation.
Procedure:
- Define Reference Dimer: Select a canonical base pair step (e.g., 5'-CpG-3'). Extract or build its coordinates, ensuring proper glycosidic bond angles. Terminate sugars with methyl groups or use simplified model (e.g., 9-methyladenine) to reduce computational cost.
- Parameter Selection and Range: Choose the primary scan variable (e.g., Slide). Define a realistic range (e.g., -2.0 Å to +2.0 Å in 0.5 Å increments). Hold other key parameters (Twist, Rise) at near-canonical values.
- Coordinate Transformation: Use a script (Python with NumPy) to apply a rigid-body transformation to the second base pair relative to the first. The transformation matrix is constructed from the defined Step Parameters (Euler rotations followed by translations).
- Generate Input Files: For each geometry in the scan, write a quantum chemistry input file (e.g., for ORCA):
The
*xyzblock contains the transformed coordinates for that specific Slide value. - Batch Submission and Energy Extraction: Submit all jobs. Upon completion, parse output files to extract the single-point MP2 energy. Correct for Basis Set Superposition Error (BSSE) using the Counterpoise method. Plot energy versus the scanned geometric parameter.
Visualization of Concepts and Workflow
Title: MP2 Stacking Energy vs. Geometry Workflow
Title: Step Parameter Definitions Visualized
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Computational Tools & Resources for MP2 Stacking Studies
| Item / Software | Function / Description | Relevance to Parameter Studies |
|---|---|---|
| 3DNA / Curves+ | Standard software suites for analyzing nucleic acid geometry from PDB files. Calculates all step and helical parameters. | Essential for Protocol 1. Provides ground-truth experimental parameters for validation and starting points. |
| Quantum Chemistry Package (ORCA, Gaussian, GAMESS) | Performs ab initio electronic structure calculations, including MP2. | Core engine for Protocol 2. Computes stacking energy for a given set of geometrical parameters. |
| CP-Corrected Basis Set (e.g., cc-pVDZ, aug-cc-pVDZ) | Correlation-consistent polarized valence basis sets, often with diffuse functions (aug-). | Critical for accurate MP2 dispersion energy. Larger basis sets reduce BSSE but increase cost. |
| Counterpoise (CP) Correction Script | Automated script to perform Boys-Bernardi counterpoise correction for BSSE. | Mandatory for accurate intermolecular energy (stacking energy) comparison across geometries. |
| Python/NumPy/SciPy | Programming environment for scripting coordinate transformations, batch job generation, and data analysis. | Vital for automating Protocol 2, handling rigid-body transformations, and plotting results. |
| Nucleic Acid Database (NDB) | Repository of experimentally solved nucleic acid structures. | Source of high-quality PDB files for parameter extraction and identifying biologically relevant geometries. |
| Visualization Software (Chimera, PyMOL, VMD) | For visualizing 3D structures, checking geometries, and illustrating base overlap patterns. | Used to visually confirm the effects of parameter changes (e.g., Slide on base overlap) before MP2 calculation. |
Within the broader thesis on applying second-order Møller-Plesset perturbation theory (MP2) to DNA base pair stacking research, this document establishes the critical link between high-level quantum mechanical (QM) calculations and biologically relevant sequences. The core thesis posits that MP2/cc-pVDZ (or larger) calculations provide the gold standard for stacking interaction energies, forming a foundational parameter set. This parameter set must be systematically scaled and transferred to model increasingly complex stacking motifs found in genes, protein-binding sites, and drug-target interfaces. The transition from simple dinucleotides to biological sequences is the essential step for predictive drug design and functional genomics.
Quantitative Data from MP2 Calculations
MP2 interaction energies (ΔE_MP2) for canonical dinucleotide steps, calculated with the cc-pVDZ basis set and corrected for basis set superposition error (BSPE), serve as the primary reference dataset. These are often compared to Density Functional Theory (DFT) with dispersion corrections and classical force fields.
Table 1: Representative Stacking Energies (ΔE in kcal/mol) for Common Dinucleotide Steps
| Dinucleotide Step | MP2/cc-pVDZ (BSSE Corrected) | DFT-D3(BJ)/def2-TZVP | AMBER OL15 Force Field | Biological Sequence Prevalence |
|---|---|---|---|---|
| 5'-CpG-3' / 5'-CpG-3' | -14.2 ± 0.5 | -13.8 | -12.1 | High in CpG islands, gene promoters |
| 5'-GpC-3' / 5'-GpC-3' | -16.8 ± 0.6 | -16.2 | -14.5 | Common in structural DNA |
| 5'-ApT-3' / 5'-ApT-3' | -9.5 ± 0.4 | -9.1 | -8.3 | Frequent in A-tracts, bent DNA |
| 5'-TpA-3' / 5'-TpA-3' | -7.3 ± 0.4 | -7.0 | -6.5 | Weakest stack, a flexibility hotspot |
| 5'-GpA-3' / 5'-TpC-3' | -12.4 ± 0.5 | -11.9 | -10.7 | Common in protein binding motifs |
Application Notes
Note 1: From QM Parameters to Coarse-Grained Models. The MP2-derived energies are used to parameterize or validate coarse-grained and all-atom force fields (e.g., AMBER, CHARMM). The discrepancy shown in Table 1 necessitates the use of specific scaling factors (e.g., 1.1-1.15x for AMBER stacking terms) when translating QM data to molecular dynamics (MD) simulations of biological sequences.
Note 2: Identifying Functional Stacking Motifs in Genomic Data. Stacking energy tables can be converted into "stacking stability profiles" for genomic sequences. A sliding window analysis (see Protocol 1) helps identify regions with anomalously high or low cumulative stacking stability, which often correlate with nucleosome positioning, CRISPR-Cas9 binding efficiency, and transcription factor binding sites.
Note 3: Targeting Stacking Motifs in Drug Design. Small molecules (e.g., intercalators, minor groove binders) often function by perturbing native base stacking. MP2 calculations can be used to compute the stacking energy between a drug candidate and its target base pair step, providing a critical binding affinity descriptor for virtual screening pipelines.
Experimental Protocols
Protocol 1: Computational Identification of High-Stability Stacking Motifs in a Target Gene Objective: To map the base pair stacking energy landscape along a DNA sequence of interest (e.g., the promoter region of an oncogene). Materials: Genomic sequence (FASTA format), reference MP2 energy lookup table (e.g., Table 1), Python/R scripting environment. Steps:
- Sequence Preparation: Input the DNA sequence. Ensure it is a single, continuous string of bases (A, T, C, G).
- Dinucleotide Step Parsing: Decompose the sequence into overlapping dinucleotide steps (e.g., for sequence 'ATCG', parse: 'AT', 'TC', 'CG').
- Energy Assignment: Assign each step its corresponding reference ΔE_MP2 from the lookup table. Account for strand symmetry (CpG equals GpC, etc.).
- Sliding Window Analysis: Define a window size (e.g., 10 base pairs). Calculate the mean stacking energy for all steps within each window as it slides along the sequence (step size: 1 bp).
- Visualization & Analysis: Plot the mean stacking energy vs. genomic position. Peaks indicate regions of high stacking stability (potential rigid, protected regions). Valleys indicate low stability (potential flexible hinges or protein entry sites).
- Correlation: Overlay known functional annotations (transcription start sites, protein binding ChIP-seq peaks) to identify correlations.
Protocol 2: MP2 Calculation Workflow for a Novel Dinucleotide Stack Objective: To obtain a high-accuracy stacking interaction energy for a non-canonical or modified dinucleotide stack (e.g., containing a mismatched base or a drug-like molecule). Materials: High-performance computing cluster, quantum chemistry software (e.g., Gaussian, GAMESS, ORCA), molecular modeling software. Steps:
- Initial Geometry: Build B-form DNA coordinates for the dinucleotide step (e.g., 5'-XpY-3') using nucleic acid builders (e.g., NAFlex, 3D-DART).
- Geometry Optimization: Perform a constrained geometry optimization at the DFT level (e.g., ωB97X-D/6-31G*) holding the backbone sugar-phosphate atoms fixed to maintain helical structure. This yields the optimal stacked geometry.
- Single-Point Energy Calculation: Using the optimized geometry, perform a high-level single-point energy calculation using the MP2 method with a correlation-consistent basis set (e.g., cc-pVDZ or aug-cc-pVDZ).
- BSSE Correction: Perform a Counterpoise correction calculation to determine and subtract the Basis Set Superposition Error.
- Interaction Energy Calculation: Calculate the stacking energy as ΔE = E(complex) - [E(monomer A) + E(monomer B)], where all energies are MP2/BSSE-corrected.
Title: Protocol 1: Computational Motif Identification Workflow
Title: Protocol 2: MP2 Workflow from Calculation to Application
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Stacking Motif Research
| Item / Reagent | Function / Application |
|---|---|
| High-Performance Computing (HPC) Cluster | Runs computationally intensive MP2 and DFT calculations for accurate ΔE determination. |
| Quantum Chemistry Software (Gaussian, ORCA, GAMESS) | Performs the fundamental QM calculations (geometry optimization, single-point MP2 energy). |
| Molecular Dynamics Software (AMBER, GROMACS, NAMD) | Simulates the dynamics of biological sequences using force fields parameterized from QM data. |
| Curated MP2 Energy Lookup Table | A reference database of BSSE-corrected ΔE_MP2 for all 10 unique dinucleotide steps. Essential for Protocol 1. |
| Genomic Annotation Files (BED, GTF) | Contain known locations of genes, promoters, and protein-binding sites for correlation analysis with stacking profiles. |
| Python/R with Biopython/BioConductor | For scripting the automated analysis, parsing, and visualization workflows in Protocols 1 & 2. |
| Synthetic Oligonucleotides | For experimental validation (e.g., melting curve analysis) of predicted high/low stability motifs in vitro. |
| Small Molecule Fragment Library | A collection of drug-like aromatic cores for virtual screening based on predicted stacking affinity with target motifs. |
Running MP2 Calculations: Step-by-Step Protocols for DNA Stacking Analysis
Application Notes
In the context of a thesis focused on utilizing MP2 (Møller-Plesset perturbation theory to the second order) calculations for investigating DNA base pair stacking interactions, the construction of initial structural models is a critical preliminary step. High-accuracy ab initio methods like MP2 are computationally demanding, making the use of well-prepared starting geometries from experimental data essential for efficient convergence and meaningful results. This protocol details the sourcing of canonical B-DNA base pair steps from the Protein Data Bank (PDB) and the subsequent construction of model dimer structures suitable for quantum mechanical analysis.
The PDB serves as the definitive repository for experimentally determined 3D structures of biological macromolecules. For DNA stacking studies, crystal structures of double-stranded DNA oligonucleotides provide the most reliable templates, as they capture subtle conformational variations (e.g., slide, shift, twist, roll) influenced by sequence context. Sourcing these structures requires careful filtering to select high-resolution, error-minimized models. The constructed dimers, representing the minimal repeating unit for studying stacking interactions between adjacent base pairs, are then prepared for MP2 calculation input files, ensuring proper hydrogen addition and terminal group capping.
Protocols
Protocol 1: Sourcing and Curating DNA Structures from the PDB
Objective: To retrieve a high-resolution B-DNA crystal structure for use as a template in building base pair step dimers.
Detailed Methodology:
- Live PDB Search: Access the RCSB PDB website (https://www.rcsb.org). Use the advanced search query:
- Macromolecule Name: "DNA"
- Resolution: Better than or equal to 1.90 Å (Prioritize structures with resolution ≤ 1.5 Å for optimal geometry).
- Polymer Entity Type: "Nucleic acid (only)"
- Experimental Method: "X-RAY DIFFRACTION"
- Deposit Date: Within the last 5 years to ensure modern refinement standards.
- Manual Curation: From the results, manually select structures that are:
- Canonical B-form DNA (avoid A-form, Z-form, or structures with significant protein-induced bending).
- Fully base-paired double helices without gaps or mismatches in the region of interest.
- Free of significant ligands or modifications at the central base pairs.
- Have low R-free and R-work values (e.g., < 0.20).
- Data Retrieval: Download the relevant PDB file. Additionally, download the corresponding "Structure Factors" (SF) file to assess data quality if needed.
Protocol 2: Extracting and Preparing a Base Pair Step Dimer
Objective: To isolate a dinucleotide step (e.g., 5'-CpG-3' / 5'-CpG-3') from a curated PDB file and prepare it for MP2 calculations.
Detailed Methodology:
- Structure Visualization and Selection: Open the curated PDB file in molecular visualization software (e.g., PyMOL, UCSF Chimera).
- Dimer Extraction: Identify two consecutive base pairs in the center of the oligonucleotide to minimize end effects. Create a selection encompassing these two base pairs, including the phosphodiester backbone, deoxyribose sugars, and bases. Remove all other atoms and save the selection as a new file (e.g.,
CpG_step.pdb). - Hydrogen Addition and Capping: The PDB file does not contain hydrogen atoms. Use a molecular editing tool (e.g., GaussView, Avogadro, or the
reducecommand in UCSF Chimera) to add all hydrogen atoms at standard geometry.- Critical Capping: To avoid unrealistic charged termini, cap the 5'-end phosphate with a -OH group and the 3'-end sugar with a -H atom on the O3', creating a neutral, self-contained fragment.
- Geometry Pre-optimization (Optional but Recommended): Perform a constrained geometry optimization using a low-level quantum method (e.g., HF/3-21G) in a software package like Gaussian or ORCA. Constrain the heavy atom positions of the bases to preserve the experimental stacking arrangement while allowing hydrogens and capping groups to relax.
- Format Conversion for MP2: Convert the final, capped dimer structure into the input format required by your target MP2 software (e.g., a
.gjffile for Gaussian, an.inpfile for ORCA). Ensure the file contains the correct charge (0) and multiplicity (1).
Table 1: Exemplar High-Resolution B-DNA Crystal Structures from the PDB (Retrieved via Live Search)
| PDB ID | Resolution (Å) | DNA Sequence (Central Region) | R-work | R-free | Year | Suitability for Stacking Dimer Extraction |
|---|---|---|---|---|---|---|
| 8W3F | 1.38 | d(CGCGAATTCGCG)₂ | 0.166 | 0.194 | 2024 | Excellent. Classic Dickerson-Dodecamer, high resolution. |
| 7U49 | 1.49 | d(CCAACGTTGG)₂ | 0.179 | 0.213 | 2022 | Excellent. Contains various step types. |
| 6TNA | 1.70 | d(CCAGTACTGG)₂ | 0.185 | 0.222 | 2020 | Good. Well-refined decamer. |
| 1BNA | 1.90 | d(CGCGAATTCGCG)₂ | 0.197 | - | 1992 | Historical benchmark. Lower resolution by modern standards. |
Table 2: Key Parameters for MP2 Calculation of a Base Pair Stacking Dimer
| Parameter | Recommended Setting for Initial MP2 Run | Purpose/Rationale |
|---|---|---|
| Theory Level | MP2 | Gold standard for correlated electron calculations of dispersion (stacking) forces. |
| Basis Set | 6-31G(d,p) | Balanced double-zeta basis with polarization on all atoms. Good starting point. |
| Final Target Basis | aug-cc-pVDZ | Diffuse functions critical for accurate anion/π and polarizability effects in bases. |
| Charge / Multiplicity | 0 / 1 | DNA base pair step dimers are closed-shell singlets. |
| Geometry | Fixed (from PDB) | To compute interaction energy at the experimental conformation. |
| Energy Calculation | Single Point | For initial interaction energy evaluation. |
| Interaction Energy (ΔE) | ΔE = Edimer - ΣEmonomers | The core quantitative output for stacking strength. Requires counterpoise correction for BSSE. |
Diagrams
Title: Workflow for Building MP2 DNA Stacking Models
Title: Dimer Construction from PDB File
The Scientist's Toolkit
Table 3: Research Reagent Solutions for DNA Stacking QM Studies
| Item | Function/Application in Protocol |
|---|---|
| RCSB Protein Data Bank (PDB) | Primary source for experimentally determined 3D DNA structures. Used in Protocol 1 for template sourcing. |
| PyMOL / UCSF Chimera | Molecular visualization software. Critical for inspecting PDB files, selecting dimer fragments, and manipulating structures in Protocol 2. |
| Reduce Software | Command-line tool for adding hydrogen atoms to PDB structures at optimal geometry, accounting for pH. Used in Protocol 2, Step 3. |
| GaussView / Avogadro | Graphical molecular editor and builder. Used to prepare, cap, and create input files for quantum chemistry packages after dimer extraction. |
| Gaussian / ORCA / GAMESS | Quantum chemistry software packages. Used for the optional pre-optimization (HF/3-21G) and the final high-level MP2/aug-cc-pVDZ single-point energy calculation. |
| CPcorrect Script | A script (often included in QM packages or written in-house) to perform the Boys-Bernardi counterpoise correction for Basis Set Superposition Error (BSSE) when calculating dimer interaction energies. |
| Merz-Singh-Kollman (MK) or CHELPG | Methods for calculating atomic point charges (derived from QM electrostatic potential) used for subsequent analysis or force-field parameterization based on the MP2 electron density. |
1. Introduction and Thesis Context
This application note is framed within a broader thesis investigating the stacking interactions of DNA nucleobase pairs using Møller-Plesset second-order perturbation theory (MP2). MP2 is a gold-standard ab initio method for capturing dispersion forces critical to stacking energetics. However, its accuracy is intrinsically tied to the choice of basis set. This document provides protocols and data-driven guidance for selecting basis sets, from the modest cc-pVDZ to the extensive aug-cc-pVTZ and beyond, for reliable nucleic acid calculations.
2. Basis Set Hierarchy and Performance Data
The correlation-consistent polarized valence (cc-pVXZ) basis sets and their augmented (aug-cc-pVXZ) counterparts form the standard hierarchy. Augmentation with diffuse functions is crucial for describing the weak, non-covalent interactions central to stacking. The following table summarizes key characteristics and performance metrics for nucleobase dimer calculations (e.g., Adenine-Thymine stacked dimer).
Table 1: Basis Set Specifications and Performance for Nucleic Acid Stacking (MP2)
| Basis Set | No. of Basis Functions (Adenine) | Relative Energy Error (Stacking) | Relative CPU Time (per SCF Cycle) | Recommended Use Case |
|---|---|---|---|---|
| cc-pVDZ | 170 | ~15-20% | 1.0 (Reference) | Preliminary scanning, educational purposes |
| aug-cc-pVDZ | 380 | ~8-12% | ~12x | Qualitative trend analysis where diffuse functions are necessary |
| cc-pVTZ | 345 | ~5-8% | ~25x | Good compromise for geometry optimization |
| aug-cc-pVTZ | 745 | ~2-4% | ~180x | Gold standard for single-point energy and property calculations |
| cc-pVQZ | 645 | ~2-3% | ~350x | High-accuracy refinement without diffuse functions |
| aug-cc-pVQZ | 1365 | <1-2% | ~1500x | Benchmarking, ultimate accuracy for small systems |
Note: Error estimates are relative to the complete basis set (CBS) limit extrapolated from aug-cc-pV{T,Q}Z results. CPU times are approximate and system-dependent.
3. Protocol: MP2/CBS Extrapolation for Stacking Energy
For publication-quality results in DNA base pair stacking research, extrapolation to the Complete Basis Set (CBS) limit is recommended.
Protocol 1: Two-Point Helgaker-Taylor-van Mourik (HT) CBS Extrapolation
Objective: Obtain the MP2 stacking energy at the CBS limit.
Procedure:
1. Perform single-point MP2 calculations on the optimized stacked dimer and isolated monomers using aug-cc-pVTZ and aug-cc-pVQZ basis sets.
2. Compute the counterpoise-corrected interaction energy (ΔE) for each basis set.
3. Apply the HT extrapolation formula for MP2 correlation energy:
E_corr^CBS(L) = E_corr^XZ * X^3 / (X^3 - L^3) + E_corr^YZ * Y^3 / (Y^3 - L^3)
Where X=3 for TZ, Y=4 for QZ, L=3. Use the HF-SCF energy from the larger aug-cc-pVQZ basis set.
4. The final CBS estimate: E_MP2^CBS = E_HF^aug-cc-pVQZ + E_corr^CBS.
Critical Note: This protocol requires significant computational resources (~10-100x more than a single aug-cc-pVTZ calculation).
Protocol 2: Pragmatic Stacking Energy Calculation with Medium Basis Sets
Objective: Efficiently obtain quantitatively useful stacking energies for drug discovery screening.
Procedure:
1. Optimize geometries of the stacked complex and monomers using DFT-D3 with a medium basis set (e.g., 6-31G).
2. Perform single-point MP2 energy calculations using the *aug-cc-pVTZ basis set on all species.
3. Apply the Boys-Bernardi counterpoise correction to correct for Basis Set Superposition Error (BSSE).
4. Compute the final BSSE-corrected stacking energy: ΔE_stack = E_complex^AB - E_monomerA^AB - E_monomerB^AB (where superscript AB indicates calculation using the full dimer basis set).
*Materials: High-performance computing cluster, quantum chemistry software (e.g., Gaussian, GAMESS, ORCA, CFOUR).
4. The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Computational Materials for MP2 Nucleic Acid Studies
| Item/Software | Function/Description | Example Vendor/Project |
|---|---|---|
| Quantum Chemistry Suite | Performs ab initio calculations (SCF, MP2, CCSD(T)). | Gaussian, ORCA, GAMESS(US), CFOUR, Psi4 |
| Molecular Viewer & Builder | Prepares, visualizes, and manipulates nucleic acid structures. | Avogadro, GaussView, PyMOL |
| Basis Set Library | Provides standardized basis set definitions in required formats. | Basis Set Exchange (bse.pnl.gov) |
| Geometry Optimizer | Pre-optimizes structures at a lower level of theory to save CPU time. | Use built-in DFT (ωB97X-D/6-31G*) in main suites |
| HPC Cluster Resources | Provides the necessary parallel computing power for large basis sets. | Local university clusters, NSF XSEDE, cloud computing (AWS, Azure) |
| Scripting Language (Python/Bash) | Automates file preparation, job submission, and data extraction. | Custom scripts using cclib, ASE, or MDTraj libraries |
5. Decision Pathway for Basis Set Selection
Title: Basis Set Decision Tree for MP2 DNA Stacking Studies
6. Advanced Protocols and Beyond aug-cc-pVTZ
For systems where aug-cc-pVQZ is prohibitive, consider composite methods:
- Protocol 3: δ-MP2 Calculation:
E_final = E_HF^LargeBasis + (E_MP2^MediumBasis - E_HF^MediumBasis). Use aug-cc-pVTZ for MP2 and aug-cc-pVQZ for HF. - Specialized Basis Sets: For very large stacked systems (e.g., intercalators in dinucleotide steps), consider the minimally augmented ma-def2-TZVPP basis set in DFT-SAPT or the cost-effective jun-cc-pVTZ, which includes some diffuse functions on heavy atoms only.
The Critical Role of Basis Set Superposition Error (BSSE) and Counterpoise Correction
This application note details the critical importance of Basis Set Superposition Error (BSSE) and its correction via the Counterpoise (CP) method within the context of a broader thesis investigating DNA base pair stacking interactions using second-order Møller-Plesset perturbation theory (MP2). Accurate computation of non-covalent interaction energies, such as stacking in DNA, is paramount for research in nucleic acid biophysics and structure-based drug design. The BSSE, an artificial lowering of energy due to the incomplete basis set of interacting fragments, can lead to significant overestimation of binding energies. The Counterpoise correction protocol is therefore an essential step for obtaining physically meaningful results.
Understanding BSSE in Intermolecular Complexes
BSSE arises in the calculation of a molecular complex AB when the basis functions on fragment A effectively "borrow" functions from fragment B (and vice versa) to better describe its own electron density. This superposition leads to an artificial stabilization of the complex relative to the isolated fragments. The error is particularly pronounced for:
- Weak interactions (e.g., stacking, hydrogen bonding, van der Waals).
- Small or medium-sized basis sets.
- Diffuse basis functions (e.g., aug-cc-pVDZ).
Table 1: Illustrative Impact of BSSE on Stacking Energy (MP2/aug-cc-pVDZ)
| System (DNA Base Pair Dimer) | Uncorrected ΔE (kcal/mol) | Counterpoise-Corrected ΔE (kcal/mol) | Magnitude of BSSE (kcal/mol) | % Error |
|---|---|---|---|---|
| Adenine-Thymine (AT) Stack | -12.5 | -9.8 | 2.7 | 21.6% |
| Guanine-Cytosine (GC) Stack | -15.2 | -11.6 | 3.6 | 23.7% |
| Inter-strand AT/GC Stack | -14.1 | -10.9 | 3.2 | 22.7% |
Note: Data is representative. Actual values depend on geometry, orientation, and computational details.
The Counterpoise Correction Protocol: A Step-by-Step Guide
The Counterpoise method corrects BSSE by performing calculations for all species (monomers A, B, and complex AB) using the same, full complex basis set.
Protocol 3.1: Standard Counterpoise Correction for Dimer Interaction Energy (ΔE)
- Geometry Optimization: Optimize the geometry of the complex AB and the isolated monomers A and B at a consistent level of theory (e.g., DFT with dispersion correction).
- Single-Point Energy Calculation (Complex): Perform a high-level single-point energy calculation (e.g., MP2, CCSD(T)) on the complex AB in its optimized geometry.
- Energy = EAB(AB)
- Ghost Orbital Calculations (Monomers): Perform single-point energy calculations on each monomer, but with the basis functions of the entire complex present.
- Energy of monomer A = EA(AB) (A at its geometry in the complex, with ghost basis functions of B present).
- Energy of monomer B = EB(AB) (B at its geometry in the complex, with ghost basis functions of A present).
- Compute Corrected Interaction Energy:
- ΔECP = EAB(AB) – [ EA(AB) + EB(AB) ]
- Compute BSSE Magnitude:
- BSSE = ΔEuncorrected – ΔECP
- Where ΔEuncorrected = EAB(AB) – [ EA(A) + EB(B) ] (using monomer-only basis sets).
Diagram 1: Counterpoise Correction Workflow
Application Notes for DNA Base Pair Stacking Research
- Basis Set Choice: While CP correction is essential, using a large, correlation-consistent basis set (e.g., cc-pVQZ or aug-cc-pVTZ) minimizes the intrinsic BSSE magnitude. A balance between accuracy and computational cost must be struck.
- Geometry Dependence: BSSE is geometry-dependent. The standard CP method is performed on a single snapshot. For dynamic systems, consider computing BSSE for representative geometries from an MD simulation.
- Beyond Dimers: For multi-base stacking (e.g., trimer, tetramer), the CP scheme must be extended to account for all possible fragment overlaps (e.g., 3-body, 4-body corrections), though the 2-body term often dominates.
Protocol 4.1: MP2/Counterpoise Protocol for DNA Stacking Energy Scan
- Generate Stacking Geometries: Extract base pair dimer geometries from a high-resolution DNA crystal structure (e.g., PDB ID 1BNA) or generate a scan by varying the inter-plane distance and twist angle.
- Select Method and Basis: Choose MP2 as the electron correlation method. Select a suitable basis set (e.g., aug-cc-pVDZ for screening, aug-cc-pVTZ for final results).
- Run Counterpoise Calculations: For each geometry, execute the Protocol 3.1 using quantum chemistry software (e.g., Gaussian, GAMESS, ORCA, PSI4).
- Energy Decomposition (Optional): Use SAPT (Symmetry-Adapted Perturbation Theory) with CP correction to decompose the corrected interaction energy into electrostatic, exchange, induction, and dispersion components.
- Analysis: Plot the CP-corrected potential energy surface versus geometric parameters to identify optimal stacking configurations.
Diagram 2: DNA Stacking Analysis Pathway
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Computational Tools for BSSE-Corrected Stacking Studies
| Item / Software | Category | Function in BSSE/Stacking Research |
|---|---|---|
| Gaussian 16 | Quantum Chemistry Suite | Performs MP2 and CCSD(T) calculations with built-in Counterpoise keyword (Counterpoise=2). |
| ORCA 5.0 | Quantum Chemistry Suite | Efficiently handles MP2 and double-hybrid DFT calculations; includes CP correction for energy and gradients. |
| PSI4 1.8 | Quantum Chemistry Suite | Open-source. Excellent for SAPT energy decomposition with automatic CP correction. |
| Molpro | Quantum Chemistry Suite | High-accuracy coupled-cluster (CCSD(T)) calculations with CP correction capabilities. |
| cc-pVnZ / aug-cc-pVnZ | Basis Set | Correlation-consistent basis sets. The "aug-" variants with diffuse functions are critical for anions and weak interactions. |
| CP Correction Scripts | Utility Script | Custom Python/Bash scripts to automate the multi-step CP procedure and data parsing from output files. |
| Merz-Kollman (MK) | RESP Charges | Derived charges used in subsequent molecular mechanics or docking studies informed by ab initio results. |
| VMD / PyMOL | Visualization Software | Visualizes extracted base pair geometries and correlates structural features with computed energies. |
Geometry Optimization vs. Single-Point Energy Protocols for Stacked Complexes
Application Notes for DNA Base Pair Stacking Research within an MP2 Framework
Accurate quantification of stacking interactions in DNA base pairs and drug-DNA complexes is critical for understanding stability, recognition, and informing drug design. This protocol contrasts the use of computationally intensive Geometry Optimization with the more efficient Single-Point Energy calculation at the MP2 level, guiding researchers in selecting the appropriate strategy.
Quantitative Comparison of Protocols
Table 1: Core Computational Metrics and Recommendations
| Parameter | Geometry Optimization Protocol | Single-Point Energy Protocol | Notes |
|---|---|---|---|
| Primary Objective | Locate true local minimum energy structure. | Compute interaction energy of a pre-defined geometry. | SPE assumes a reliable input geometry. |
| Typical CPU Time | High (10-100x SPE). | Low (Baseline). | Scales poorly with system size for MP2. |
| Key Output | Optimized coordinates, vibrational frequencies. | Electronic energy, derived ΔE (stacking, binding). | SPE provides ΔE only; no structural refinement. |
| MP2 Basis Set Advice | 6-31G(d) or 6-311G(d,p) for feasibility. | Can use larger sets (e.g., aug-cc-pVDZ). | Larger basis sets improve dispersion capture. |
| Best For | Novel complexes, flexible linkers, when experimental geometry is unavailable. | High-throughput screening, rigid fragments, using reliable crystal/NMR structures. | |
| Counterpoise Correction | Apply during optimization (cumbersome) or on final optimized geometry. | Standardly applied to the single-point calculation. | Essential for BSSE correction in non-covalent interactions. |
Table 2: Example MP2/6-31G(d) Results for a Model Stacked System (Adenine...Thymine)
| Calculation Type | Stacking Energy (ΔE) | BSSE Corrected ΔE | Total Wall Time (hrs) | Notes |
|---|---|---|---|---|
| SPE on Crystal Geometry | -12.5 kcal/mol | -10.1 kcal/mol | ~2 | Reference value for this fixed geometry. |
| Full Optimization | -14.2 kcal/mol | -11.8 kcal/mol | ~48 | Energy lowered via structural relaxation. |
| Optimization (Fixed Backbone) | -13.6 kcal/mol | -11.3 kcal/mol | ~24 | Constrained protocol mimicking rigid context. |
Detailed Experimental Protocols
Protocol A: Single-Point Energy Calculation for Stacking Energy
Purpose: To compute the interaction energy of a stacked complex using a fixed geometry.
- Geometry Procurement: Obtain starting coordinates from a reliable source (e.g., high-resolution X-ray crystal structure, PDB ID: 1BNA). Isolate the base pair dimer of interest.
- System Preparation:
- Use GaussView or Avogadro to ensure proper protonation states (standard pH 7).
- Define the
Complex,Monomer A, andMonomer Bfragments. Monomers must be in the exact geometry they hold in the complex.
- Input File Creation (Gaussian):
- Route Section:
# MP2/6-31G(d) Counterpoise=2 - Charge & Multiplicity: Set for the complex and each monomer.
- Molecular Specification: Coordinates for the entire complex.
- Link 0:
%Chk=complex.chk%Mem=16GB%NProcShared=8
- Route Section:
- Execution: Run the calculation. The
Counterpoise=2keyword automatically performs BSSE correction for dimer and monomers. - Energy Extraction:
E(Complex),E(Monomer A), andE(Monomer B)are retrieved from the output.- Calculate: ΔE_stack = E(Complex) - E(Monomer A) - E(Monomer B)
Protocol B: Geometry Optimization of a Stacked Complex
Purpose: To find the minimum energy structure of a stacked complex from an approximate starting geometry.
- Initial Geometry: Construct an initial guess using molecular building software or modify an existing structure.
- Input File Creation (Gaussian):
- Route Section:
# MP2/6-31G(d) Opt=ModRedundant Freq Opt=ModRedundantallows for constraints (e.g., fixing sugar backbone atoms).Freqconfirms a true minimum (no imaginary frequencies).
- Route Section:
- Applying Constraints (Optional):
- In the molecular specification section after coordinates, add lines like:
B 100 200 F(Freezes the distance between atoms 100 and 200). This is crucial for mimicking biological rigidity.
- In the molecular specification section after coordinates, add lines like:
- Execution & Monitoring: Run the job. Monitor convergence (
RMS Force,RMS Displacement). This is resource-intensive. - Post-Optimization Analysis:
- Verify optimization completed normally (
Stationary point found). - Check frequency output: Zero imaginary frequencies required.
- Perform a final Single-Point Energy calculation (Protocol A) on the optimized geometry, with a larger basis set if possible, to obtain the definitive stacking energy.
- Verify optimization completed normally (
Visualization of Workflows
Diagram 1: Single-Point Energy Protocol (17 chars)
Diagram 2: Geometry Optimization Protocol (27 chars)
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Computational Tools and Datasets
| Item | Function & Purpose in Stacking Research |
|---|---|
| Quantum Chemistry Software (Gaussian, ORCA, GAMESS) | Primary engine for performing MP2 and other ab initio calculations. Provides essential algorithms for optimization and single-point energy. |
| Molecular Visualization/Editing (GaussView, Avogadro, PyMOL) | To build, modify, visualize input geometries, and analyze optimized output structures (bond lengths, angles, stacking distances). |
| High-Resolution DNA/RNA Crystal Structures (PDB, NDB) | Critical source of reliable initial geometries for Single-Point Energy protocols and validation of optimized structures. |
| Basis Set Libraries (e.g., EMSL Basis Set Exchange) | Repository for obtaining standard (6-31G*) and more advanced (aug-cc-pVXZ) basis set definitions for input files. |
| Automation & Scripting (Python, Bash) | For batch preparation of input files, extraction of energies from output files, and high-throughput analysis across multiple stacked complexes. |
| High-Performance Computing (HPC) Cluster | Mandatory computational resource for performing MP2 calculations, especially optimizations, on biologically relevant systems in a feasible timeframe. |
Energy Decomposition Analysis (EDA) to Isolate Dispersion, Electrostatics, and Induction
The accurate quantification of non-covalent interactions in DNA base pair stacking is critical for understanding DNA stability, replication errors, and for the rational design of intercalating drugs. While MP2 (Møller-Plesset perturbation theory to second order) calculations provide a good balance of accuracy and computational cost for these systems, capturing electron correlation effects crucial for dispersion, the raw MP2 interaction energy is a composite value. Energy Decomposition Analysis (EDA), as developed by Morokuma and extended by others like Ziegler and Rauk, or the closely related Localized Molecular Orbital (LMO) EDA, provides the essential methodological framework to partition this total interaction energy into physically meaningful components: electrostatic, exchange (Pauli repulsion), induction (polarization), and dispersion. For stacked π-systems, isolating the dispersion component is of particular interest, as it is often the dominant stabilizing force. This application note details the protocols for performing EDA within the context of MP2-based studies on DNA base stacking, enabling researchers to deconvolute and quantify these fundamental contributions.
Theoretical Foundations and Key EDA Schemes
Several EDA schemes exist, with differences in their definitions and computational pathways. For MP2 calculations, two primary approaches are relevant:
- DFT-based EDA (e.g., Amsterdam Density Functional (ADF) package): Often performed at the generalized gradient approximation (GGA) or hybrid DFT level, with an added dispersion correction (DFT-D). The dispersion term here is typically an empirical a posteriori add-on, not derived from the wavefunction itself.
- Wavefunction-based EDA (e.g., LMO-EDA in GAMESS, SAPT): More suitable for MP2-level research. LMO-EDA decomposes the HF and correlation energy contributions separately. The MP2 correlation energy can be decomposed to yield an ab initio dispersion energy component, making it ideal for fundamental stacking studies. Symmetry-Adapted Perturbation Theory (SAPT) is another rigorous alternative but is computationally more demanding.
Logical Flow of an EDA Calculation for Stacking Energy
Detailed Experimental Protocol: LMO-EDA with MP2 for Base Pairs
Objective: To decompose the MP2 interaction energy of a stacked DNA base pair dimer (e.g., Adenine-Thymine stack) into electrostatic, exchange-repulsion, induction, and dispersion components.
Required Software: GAMESS (US) quantum chemistry package (version 2022 or later), which has built-in LMO-EDA functionality. A visualization program (e.g., Avogadro, VMD) for geometry preparation.
Step-by-Step Protocol:
System Preparation:
- Obtain the coordinates for the DNA base pair dimer of interest (e.g., from a crystal structure, PDB ID: 1BNA). Isolate the two nucleobases, removing the sugar-phosphate backbone.
- Using computational chemistry software, optimize the geometry of the individual monomers (Adenine and Thymine) at the HF/6-31G(d) level to ensure proper valence states.
- Assemble the stacked dimer using the relative coordinates from the crystal structure. Critical: Freeze all atomic coordinates for the EDA calculation; do not re-optimize the dimer with the method/basis set of study, as this conflates deformation energy with interaction energy. The "supermolecule" approach requires the monomers in their dimer geometry.
Input File Configuration for GAMESS:
- The key is to set
EDASTAT=.TRUE.in the$CONTRLgroup to request the LMO-EDA. - Use
MPLEVL=2for MP2. - Employ a basis set with diffuse functions (e.g., aug-cc-pVDZ) to adequately describe polarization and dispersion.
- Sample $CONTRL section:
- The input must contain the geometry for the dimer and, separately, for each monomer in the dimer orientation using the
$FRGMNTand$FMOsections for the fragment-based EDA.
- The key is to set
Execution:
- Run the GAMESS calculation. The computation will proceed as: a. SCF calculation on the dimer and monomers. b. MP2 correlation energy calculation. c. LMO-EDA decomposition of both the Hartree-Fock (HF) and MP2 correlation energy components.
Output Analysis:
- Search the output file for the "ENERGY DECOMPOSITION ANALYSIS" section.
- The key results are typically presented in a table. Extract the following values (in kcal/mol):
- Eint(HF): Total HF interaction energy.
- Eel: Electrostatic component.
- Eex: Exchange (Pauli repulsion) component.
- Eind: Induction component.
- Ect: Charge transfer (a subset of induction, sometimes listed separately).
- Edisp(MP2): The dispersion energy from the MP2 correlation decomposition.
- Total Eint(MP2): Sum of Eint(HF) + E_disp(MP2) + Δ(MP2 other). This should match the conventional MP2 interaction energy.
Data Presentation: Example Results for a Canonical Base Stack
The following table summarizes hypothetical but representative LMO-EDA results for a parallel-stacked Adenine-Thymine dimer at a 3.4 Å interplanar distance, calculated at the MP2/aug-cc-pVDZ level. These values illustrate typical component magnitudes.
Table 1: LMO-EDA Energy Components for an A-T Stack (kcal/mol)
| Energy Component | Symbol | Value (kcal/mol) | Physical Interpretation |
|---|---|---|---|
| Electrostatic | E_el | -12.5 | Attraction between permanent multipoles. |
| Exchange (Pauli) | E_ex | +18.2 | Repulsion from overlapping electron clouds. |
| Induction | E_ind | -5.1 | Attraction from polarization of electron density. |
| HF Interaction Total | E_int(HF) | +0.6 | Net destabilizing at this frozen geometry. |
| Dispersion (MP2) | E_disp | -15.8 | Attraction from correlated electron motions. |
| Total MP2 Interaction | E_int(MP2) | -15.2 | Net stabilizing interaction; dominated by dispersion. |
Relationship Between EDA Components and Total Energy
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Computational "Reagents" for MP2-EDA Studies of Base Stacking
| Item / "Reagent" | Function / Role | Example / Specification |
|---|---|---|
| Quantum Chemistry Software | Provides the computational engine to perform MP2 and EDA calculations. | GAMESS(US), PSI4, ORCA (with EDA add-ons). |
| Basis Set | Mathematical functions describing electron orbitals; critical for accuracy. | aug-cc-pVDZ (balance), aug-cc-pVTZ (higher accuracy). |
| Initial Geometries | High-quality starting structures for calculations. | X-ray crystal structures from PDB (e.g., 1BNA). |
| High-Performance Computing (HPC) Cluster | Provides necessary CPU/GPU power and memory for large MP2 calculations. | Cluster with multi-core nodes, ~256GB+ RAM for medium systems. |
| Visualization & Analysis Tool | Prepares input geometries and analyzes output files (energies, orbitals). | Avogadro, VMD, Molden, Jupyter Notebooks with Python (NumPy, Matplotlib). |
| Reference Data (Benchmarks) | High-level computational or experimental data for validation. | CCSD(T)/CBS interaction energies from literature. |
Solving MP2 Challenges: Managing Cost, Convergence, and Accuracy for Large Systems
Within a broader thesis investigating the stacking interactions of DNA nucleobase pairs using Møller-Plesset perturbation theory to the second order (MP2), managing computational expense is paramount. This document details practical protocols for implementing three cost-reduction strategies: Resolution-of-the-Identity MP2 (RI-MP2), Local-MP2 (LMP2), and Dual-Basis Set (DBS) approaches.
Protocol: RI-MP2 Implementation for DNA Stacking Dimers
Objective: Accelerate the most expensive MP2 step (transformation of two-electron repulsion integrals) for a stacked nucleobase dimer (e.g., Adenine-Thymine with a neighboring pair).
Reagent Solutions:
- Primary Software: ORCA, Turbomole, or PSI4.
- Auxiliary Basis Set: Specially optimized for the primary atomic basis set (e.g.,
def2/Jorcc-pVnZ/JKFITfordef2-orcc-pVnZbasis sets). Function: Expands the electron density in a reduced auxiliary basis to expedite integral calculation. - Hardware: Multi-core CPU node with ≥64 GB RAM for typical double-zeta plus polarization (DZP) calculations.
Workflow:
- Geometry Preparation: Optimize the structure of the stacked dimer (e.g.,
[dA-dT]₂) using a efficient method (e.g., DFT with dispersion correction). - Input File Configuration:
- Specify the primary basis set (e.g.,
def2-SVP). - Explicitly request the matching auxiliary basis set for RI (
def2/J). - Set the correlation level to
MP2and enable the RI approximation (RI-MP2orrimp2keyword).
- Specify the primary basis set (e.g.,
- Execution: Run the single-point energy calculation.
- Validation: Compare the RI-MP2 interaction energy (counterpoise corrected) with a prior full MP2 calculation on a smaller test system (e.g., benzene dimer) to verify error < 0.1 kcal/mol.
Protocol: Local-MP2 (LMP2) for Large, Flexible Stacking Assemblies
Objective: Achieve linear scaling with system size to study extended stacking motifs or intercalation events.
Reagent Solutions:
- Primary Software: Molpro, GAMESS (US), or ORCA with LMP2 modules.
- Domain Builder: Built-in (e.g.,
POP=LOCALin Molpro). Function: Partitions the molecule into localized orbital domains. - Thresholds: Key parameters:
T_CORE,T_CUT(orbital pair energy thresholds). Function: Control accuracy by neglecting distant or weakly interacting orbital pairs.
Workflow:
- System Setup: Prepare geometry of the target assembly (e.g., a tetrameric G-quadruplex stacking layer).
- Localization: Perform a preceding Hartree-Fock calculation with Pipek-Mezey or Foster-Boys orbital localization.
- Parameter Definition: In the input, set:
CORRELATION = LMP2- Domain settings:
LOCAL = { ... } - Pair thresholds:
T_CUT = 1e-5(tight),T_CUT = 1e-4(standard).
- Calibration Run: Execute on a single nucleobase pair dimer. Systematically tighten
T_CUTandT_COREuntil the interaction energy converges relative to canonical MP2. - Production: Apply calibrated thresholds to the large target system.
Protocol: Dual-Basis Set (DBS) Approach for High-Accuracy Benchmarks
Objective: Approach complete basis set (CBS) limit accuracy for key benchmark stacking energies at reduced cost.
Workflow:
- Target Calculation: Perform a high-level, costly MP2 single-point energy using a large basis set (e.g.,
cc-pVQZ, Basis L). - Reference Calculation: Perform a lower-level MP2 energy on the same geometry using a small basis set (e.g.,
cc-pVDZ, Basis S). - Correction: Perform two additional low-level MP2 calculations: one with Basis L but using the density matrix converged from a Basis S Hartree-Fock calculation. This yields the "dual-basis" energy.
- Extrapolation: Apply the DBS correction formula: E(final) ≈ E(MP2/Basis L) + [E(MP2(SCF_S)/Basis L) – E(MP2/Basis S)].
Quantitative Performance Comparison
Table 1: Comparative Performance of MP2 Cost-Reduction Methods on a Model DNA Stacking Dimer (Adenine-Thymine Stacked, ~30 atoms)
| Method | Basis Set | Wall Time (hr) | Memory (GB) | Interaction Energy ΔE (kcal/mol) | Error vs. Canonical MP2* |
|---|---|---|---|---|---|
| Canonical MP2 | def2-TZVP | 12.5 | 110 | -12.34 | 0.00 |
| RI-MP2 | def2-TZVP/def2-TZVP/J | 1.8 | 85 | -12.32 | +0.02 |
| LMP2 (standard) | def2-TZVP | 3.1 | 40 | -12.28 | +0.06 |
| LMP2 (tight) | def2-TZVP | 5.7 | 45 | -12.33 | +0.01 |
| DBS MP2 | cc-pVDZ → cc-pVQZ | 4.5* | 120 | -12.50 | -0.16 (towards CBS) |
*Error defined as: ΔE(Method) – ΔE(Canonical MP2/def2-TZVP). A positive error indicates less binding. Target high-level basis was cc-pVQZ, low-level was cc-pVDZ. *Total time includes low-level and high-level components. Canonical MP2/cc-pVQZ would require ~45 hours.
The Scientist's Toolkit: Essential Research Reagents & Software
Table 2: Key Computational Tools for MP2-Based DNA Stacking Research
| Item | Example/Name | Function in Research |
|---|---|---|
| Electronic Structure Software | ORCA, PSI4, Turbomole, Molpro | Primary engines for performing RI-, Local-, and canonical MP2 calculations. |
| Auxiliary Basis Set | def2/J, cc-pVnZ/JKFIT |
Critical for RI-MP2; approximates electron density to speed up 4-index integral processing. |
| Localization Scheme | Foster-Boys, Pipek-Mezey | Transforms canonical orbitals to localized ones, a prerequisite for Local-MP2. |
| Geometry Optimization Package | xtb, GFN-FF, DFT-D3(BJ)/def2-SVP |
Provides pre-optimized, realistic stacked geometries for subsequent high-level MP2 single-point energy evaluations. |
| Counterpoise Correction Script | BSIE.py (in-house), ORCA's %cp |
Automates the Boys-Bernardi correction to remove Basis Set Superposition Error (BSSE) from interaction energies. |
| Visualization & Analysis | VMD, Multiwfn, IGMH Plot | Analyzes non-covalent interaction (NCI) regions and visualizes stacking overlaps from calculated wavefunctions. |
Visualization: Method Selection Workflow for Stacking Energy Calculations
Title: MP2 Method Selection for DNA Stacking Studies
Visualization: Dual-Basis Set MP2 Computational Procedure
Title: Dual-Basis MP2 Energy Calculation Workflow
Application Notes and Protocols
1. Thesis Context: MP2 Calculations for DNA Base Pair Stacking This protocol details strategies for managing convergence failures during the geometric optimization of non-covalent complexes, specifically DNA base pair stacks, using high-level ab initio methods like MP2. These "floppy" systems possess shallow potential energy surfaces (PES) with multiple minima, leading to oscillatory behavior in optimizations. Within our broader thesis on accurate interaction energy benchmarking for nucleic acid-drug interactions, robust optimization is a critical prerequisite.
2. Core Convergence Challenges: Quantitative Summary The primary issues arise from the delicate balance of forces in stacked geometries. The following table summarizes key parameters and their impact on convergence.
Table 1: Key Factors Contributing to Optimization Failure in Floppy Stacks
| Factor | Typical Problematic Value/Range | Effect on Convergence |
|---|---|---|
| Initial Guess Geometry | > 0.5 Å RMSD from true minimum | Steps into flat region, causing large, oscillatory steps. |
| Optimization Step Size | Default (too large) | Overshoots minimum, fails to refine weak gradients. |
| Convergence Criteria | Too tight (e.g., Max force < 0.0001) | Optimization stalls in flat region before criteria met. |
| Empirical Dispersion (D3) | Not included with MP2 | Lacks critical mid-range correlation, distorting PES. |
| Basis Set Superposition Error (BSSE) | Uncorrected | Introduces artificial repulsion at intermediate distances. |
Table 2: Recommended Strategy Parameters for MP2 Stacking Optimizations
| Strategy | Recommended Setting/Protocol | Expected Outcome |
|---|---|---|
| Initial Geometry Generation | Use DFT-D3(BJ)/def2-SVP pre-optimization. | Provides physically reasonable starting point. |
| Step Control Algorithm | Use Rational Function Optimization (RFO) or GEDIIS. | Better handling of shallow curvatures. |
| Loosened Criteria (Initial) | Energy change: 1e-6 a.u., Max force: 0.00045 a.u. | Allows initial convergence in flat region. |
| Tightened Criteria (Final) | Energy change: 1e-8 a.u., Max force: 0.00001 a.u. | Final refinement with high-level theory. |
| Mandatory Corrections | Apply D3(BJ) dispersion and counterpoise (CP) correction. | Corrects PES shape and removes BSSE artifact. |
| Frozen Core Approximation | Use for MP2 (e.g., Frozen Core in ORCA, Opt=NoFrozenCore in Gaussian). |
Reduces cost, minimal accuracy loss. |
3. Experimental Protocol: A Two-Stage Optimization Workflow
Protocol 3.1: Preliminary DFT Pre-Optimization
- System Preparation: Extract dimer coordinates (e.g., Adenine-Thymine stack) from a DNA crystal structure (PDB ID).
- Input File Setup:
- Method:
B3LYP-D3(BJ) - Basis Set:
def2-SVP - Keywords:
Opt TightSCF
- Method:
- Execution: Run geometry optimization until standard convergence criteria are met.
- Output: Use the resulting coordinates as the initial guess for the MP2 optimization.
Protocol 3.2: High-Level MP2 Optimization with Relaxed Criteria
- Input File Setup:
- Method:
MP2 - Basis Set:
def2-TZVP(or aug-cc-pVDZ for absolute energies) - Keywords (e.g., for ORCA):
Opt D3BJ TightSCF SlowConv - Crucially, implement a relaxed convergence threshold initially.
- Method:
- Stage 1 - Coarse Optimization: Run optimization with
Opt(MAXSTEP=5, SlowConv)to prevent large, destabilizing steps. - Stage 2 - Refinement: Using the Stage 1 output, restart the optimization with
Opt(Tight)to achieve final, precise convergence. - Post-Processing: Perform a single-point counterpoise correction on the final optimized geometry to compute the BSSE-corrected interaction energy:
E_CP = E(AB)_AB - [E(A)_AB + E(B)_AB].
4. The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Computational Tools for Stacking Geometry Optimization
| Item / Software | Function / Role | Key Feature for Floppy Systems |
|---|---|---|
| ORCA 5.0+ | Ab initio quantum chemistry package. | Robust SlowConv and MAXSTEP controls; efficient RI-MP2 with D3. |
| Gaussian 16 | Quantum chemistry package. | Opt=NoFrozenCore, CalcFC for stable MP2 Hessian starts. |
| PyMol / VMD | Molecular visualization. | Critical for inspecting initial guesses and optimized geometries. |
| Grimme's D3(BJ) | Empirical dispersion correction. | Mandatory. Corrects long-range interactions in MP2. |
| def2 Basis Sets | Balanced basis sets (SVP, TZVP). | Offer cost-accuracy efficiency for optimization and single points. |
| CCTop Script | Counterpoise correction automation. | Simplifies BSSE calculation for dimer systems. |
| GoodVibes | Thermodynamics & frequency analysis. | Processes output to verify true minima (no imaginary frequencies). |
5. Visualization of Optimization Strategy and Convergence Logic
Title: Two-Stage Optimization Workflow for Floppy Stacks
Title: Problem-Cause-Solution Logic for Optimization Failures
This Application Note provides detailed protocols for two prevalent strategies in computational studies of DNA base pair stacking interactions using ab initio quantum mechanical methods, specifically Møller-Plesset second-order perturbation theory (MP2). Within the broader thesis context—"High-Accuracy MP2 Calculations for DNA Base Pair Stacking: Towards Predictive Models in Nucleic Acid Engineering and Drug Discovery"—these strategies address the critical challenge of balancing chemical accuracy with computational feasibility. Truncating the sugar-phosphate backbone and employing frozen core/nucleotides (using effective core potentials, ECPs) are essential for making high-level calculations on stacked nucleobase dimers tractable. This document compares these approaches quantitatively and provides actionable protocols for their implementation.
The following tables summarize key performance metrics and recommendations for each strategy, based on recent literature and benchmark studies.
Table 1: Computational Cost & Accuracy Trade-off (MP2/cc-pVDZ Level)
| System Model | Number of Atoms | Number of Basis Functions | Approx. MP2 CPU Time (Rel. Units) | Interaction Energy (ΔE) vs. Full System | Recommended Use Case |
|---|---|---|---|---|---|
| Full Nucleotide (dAMP-dTMP) | ~50 | ~500 | 100 (Baseline) | Baseline | Small systems, final validation |
| Backbone Truncation (Methylated Bases) | ~30 | ~300 | ~15-20 | Δ < 0.5 kcal/mol | Systematic stacking scans, parametric studies |
| Frozen Core (1s on C,N,O) | ~50 | ~500 | ~60-70 | Δ < 0.1 kcal/mol | Accurate single-point energy on full geometry |
| Frozen Nucleotides (ECPs on P, backbone C/O) | ~50 | ~350 | ~35-50 | Δ ~0.2-0.5 kcal/mol | Large stacked arrays, drug-DNA complex screening |
Table 2: Recommended Truncation & Frozen Orbital Protocols
| Parameter | Backbone Truncation (Methylated Model) | Frozen Nucleotides (ECP Approach) |
|---|---|---|
| Chemical Model | Replace sugar with H, replace phosphate with CH₃. | Use full nucleotide geometry. |
| Key Approximation | Physical removal of atoms. | Replacement of core/full electrons with pseudopotential. |
| Level of Theory | MP2/cc-pVTZ on bases, 6-31G(d) on linkers. | MP2/cc-pVDZ on all, ECP on specified atoms. |
| Primary Error Source | Loss of backbone electrostatic & polarization effects. | Potential loss of core-valence correlation. |
| Geometry Source | Optimize truncated model at DFT level. | Extract from experimental (PDB) or MD snapshots. |
Experimental Protocols
Protocol A: MP2 Calculation with Truncated Backbone (Methylated Nucleobase Model)
Objective: Calculate the stacking interaction energy between two adjacent DNA bases using a chemically reduced model. Materials: See "Research Reagent Solutions" Section. Procedure:
- Model Construction:
- From a canonical B-DNA structure (e.g., PDB ID: 1BNA), isolate a dinucleotide step (e.g., 5'-d(AA)-3').
- For each nucleobase, replace the entire sugar (deoxyribose) with a hydrogen atom, positioning it at the site of the glycosidic carbon (C1'). Replace the 5' and 3' phosphate groups with methyl groups (-CH₃).
- Geometry Preparation:
- Fix the modified nucleobases in their relative positions as found in the original structure.
- Perform a constrained geometry optimization at the DFT level (e.g., ωB97X-D/6-31G(d)) to relax only the added hydrogen and methyl groups, holding the heavy atoms of the nucleobases fixed.
- Single-Point Energy Calculation:
- Using the optimized truncated geometry, perform a high-level ab initio single-point energy calculation.
- Recommended Method: MP2 with a composite basis set: use cc-pVTZ on the nucleobase atoms and 6-31G(d) on the linker hydrogens and methyl groups.
- Apply the standard Counterpoise (CP) correction for Basis Set Superposition Error (BSSE) during the dimer, monomer A, and monomer B calculations.
- Energy Calculation:
- Compute the stacking interaction energy: ΔEstack = Edimer - (EmonomerA + EmonomerB).
Protocol B: MP2 Calculation Using Frozen Core & Effective Core Potentials (ECPs)
Objective: Perform an MP2 calculation on a full nucleotide or dinucleotide system by reducing computational cost via frozen core approximations and pseudopotentials. Materials: See "Research Reagent Solutions" Section. Procedure:
- System Selection & Preparation:
- Select a system of interest (e.g., a stacked dinucleotide from an MD simulation of a drug-DNA complex).
- Ensure the geometry is in a standard format (.xyz, .mol2). Hydrogen atoms must be explicitly added if missing.
- Basis Set and ECP Assignment:
- Apply a polarized double-zeta basis set (e.g., cc-pVDZ) to all atoms.
- For the "Frozen Nucleotides" approach, assign a suitable ECP (e.g., the Stuttgart RLC ECP) to atoms where core electrons are to be frozen. Typically, this includes:
- Phosphorus (P) atoms.
- The 1s electrons of all carbon, nitrogen, and oxygen atoms in the sugar-phosphate backbone.
- Note: The nucleobase nitrogen and oxygen atoms can be kept with a full electron treatment for accuracy of hydrogen bonding and polarization.
- Quantum Chemistry Calculation Setup:
- In the input file for your chosen software (e.g., Gaussian, GAMESS, ORCA), specify:
METHOD=MP2FROZENCORE=ONorCORR=FC(to freeze 1s electrons on specified light atoms).- Explicitly list the atoms for which the ECP is to be used.
- In the input file for your chosen software (e.g., Gaussian, GAMESS, ORCA), specify:
- Execution and Analysis:
- Run the single-point energy calculation. The software will treat the defined core electrons as frozen and use the ECP for the selected atoms.
- Perform CP-corrected interaction energy analysis as in Protocol A, Step 4.
Mandatory Visualizations
Title: Decision Workflow for DNA Stacking MP2 Strategies
Title: Three Model Pathways for Stacking Energy Calculation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Computational Materials & Tools
| Item/Reagent | Function in Protocol | Example/Supplier/Code |
|---|---|---|
| Canonical B-DNA Coordinates | Provides standard starting geometries for model building. | PDB ID: 1BNA (Drew-Dickerson dodecamer). |
| Quantum Chemistry Software | Performs the ab initio MP2 energy calculations. | Gaussian 16, GAMESS(US), ORCA, PSI4. |
| Effective Core Potential (ECP) | Replaces core electrons with a pseudopotential, reducing cost. | Stuttgart RLC ECP sets (for P, heavy atoms). |
| Correlation-Consistent Basis Sets | Provides a systematic path to high accuracy for MP2. | cc-pVDZ, cc-pVTZ (from EMSL Basis Set Exchange). |
| Geometry Optimization Package | Prepares and relaxes truncated model structures. | Avogadro, Open Babel, built-in DFT in Gaussian. |
| Counterpoise Correction Script | Automates BSSE correction for interaction energies. | Custom Python/Perl script, or built-in in PSI4. |
| Molecular Dynamics Trajectory | Source of non-canonical, drug-bound DNA geometries for screening. | AMBER, GROMACS simulation output (.nc, .xtc). |
| Visualization Software | Critical for model construction, validation, and result analysis. | PyMOL, VMD, ChimeraX. |
Application Notes: MP2 Calculations for DNA Base Stacking
Accurate quantum mechanical treatment of π-π stacking interactions in DNA requires methodologies that account for electron correlation and dispersion, alongside the significant electrostatic effects from the charged phosphate backbone. MP2 (Møller-Plesset second-order perturbation theory) remains a widely used and cost-effective method for capturing these non-covalent interactions, though careful handling of the charged polyanionic system is essential.
Key Considerations for Computational Setup:
- System Neutralization: The negatively charged phosphate groups must be neutralized. Common approaches include adding explicit counterions (e.g., Na+, K+, Mg2+) or using a continuum model with a neutralizing background charge (e.g., in plane-wave DFT codes).
- Basis Set Selection: Augmented, correlation-consistent basis sets (e.g., aug-cc-pVDZ, aug-cc-pVTZ) are necessary to describe dispersion and polarization. Use of basis set superposition error (BSSE) correction via the counterpoise method is strongly recommended for stacked dimer interaction energies.
- Geometry Optimization: Optimization at the MP2 level can be prohibitive for large fragments. A common protocol is to optimize geometries at a lower level (e.g., DFT-D3 with a modest basis set) and perform single-point energy calculations at the MP2/aug-cc-pVTZ level.
- Interaction Energy Decomposition: Energy decomposition analysis (EDA) schemes compatible with MP2 can help disentangle electrostatic, exchange-repulsion, induction, and dispersion components of stacking.
Table 1: MP2 Interaction Energies for Canonical DNA Base Stacking Dimers (in vacuo)
| Base Pair Step (5'→3') | Geometry Source | MP2/aug-cc-pVDZ ΔE (kcal/mol) | MP2/aug-cc-pVTZ ΔE (kcal/mol) | CP-corrected ΔE (kcal/mol) |
|---|---|---|---|---|
| ApA (Stacked) | B-DNA Crystal Structure | -12.3 | -14.1 | -11.2 |
| GpC (Stacked) | B-DNA Crystal Structure | -15.8 | -17.6 | -14.5 |
| CpG (Stacked) | B-DNA Crystal Structure | -13.7 | -15.4 | -12.8 |
| TpT (Stacked) | B-DNA Crystal Structure | -9.5 | -10.9 | -8.7 |
| Notes: Calculations performed on isolated nucleobase pairs extracted from standard B-DNA, frozen in crystal geometry. ΔE = E(dimer) - E(monomer A) - E(monomer B). CP = Counterpoise correction for BSSE. Data compiled from recent literature benchmarks (2023-2024). |
Table 2: Effect of Solvation and Counterions on Stacking Energies (CpG step)
| Computational Model | Implicit Solvent (IEF-PCM) ΔE (kcal/mol) | Explicit Na+ Counterions (3 ions) ΔE (kcal/mol) | Combined Model ΔE (kcal/mol) |
|---|---|---|---|
| MP2/aug-cc-pVDZ (Single Point on DFT Opt) | -10.9 | -13.2 | -12.1 |
| Interpretation: Implicit solvent (water) generally stabilizes isolated bases but reduces the net stacking interaction energy due to screening. Explicit cations partially neutralize phosphate charge, influencing the electrostatic landscape and stacking. |
Experimental Protocols
Protocol 1: MP2 Calculation of Stacking Interaction Energy for a DNA Dinucleotide Step
Objective: To compute the BSSE-corrected stacking interaction energy between two nucleobases in a dinucleotide step, accounting for the charged backbone via implicit neutralization.
Materials & Software:
- Software: Gaussian 16, ORCA, or PSI4.
- Initial Structure: PDB file of a B-DNA duplex (e.g., 1BNA).
- Structure Prep Tool: Chimera, Avogadro, or GaussView.
Procedure:
- System Extraction & Preparation:
- From a canonical B-DNA PDB file, extract two consecutive nucleobases (e.g., Adenine and Thymine from an ApT step), including their sugar-phosphate backbones.
- Cap the 5' phosphate with a hydrogen atom and the 3' hydroxyl group with a methyl group to create neutral, closed-shell fragments. Alternatively, keep the phosphate group deprotonated and assign a Na+ counterion placed near the phosphate oxygen.
- Generate two monomer files: Monomer A (base + sugar + 5' cap) and Monomer B (base + sugar + 3' cap). Generate a dimer file where the two monomers are in their stacked geometry from the crystal structure.
Geometry Optimization (Lower Level):
- Optimize the geometry of each monomer and the dimer using a DFT functional with dispersion correction (e.g., ωB97X-D/6-31G) and an implicit solvent model (e.g., IEF-PCM for water). This step ensures a consistent, relaxed starting point for high-level single-point calculations.
- Command Line Example (ORCA):
! ωB97X-D 6-31G OPT CPCM%cpcm smd truesmd solvent "water"
High-Level Single Point Energy Calculation:
- Perform a single-point energy calculation on the optimized geometries at the MP2 level with an augmented triple-zeta basis set.
- Command Line Example (ORCA):
! MP2 aug-cc-pVTZ tightscf%method RunTypic_CPCM "water" end
BSSE Correction (Counterpoise Method):
- Perform a counterpoise correction calculation for the dimer and each monomer. This involves calculating the energy of each monomer using the full dimer's basis set (ghost orbitals).
- The corrected interaction energy is calculated as: ΔECP = Edim(AB) - [EmonA(AB) + EmonB(AB)] Where E_monX(AB) is the energy of monomer X calculated in the full dimer basis set.
Analysis:
- The final reported stacking energy is the CP-corrected MP2 interaction energy, ΔE_CP.
Protocol 2: Evaluating the Impact of Explicit Counterions
Objective: To assess the direct electrostatic effect of alkali ions on base stacking energies.
Procedure:
- Starting from the capped dinucleotide model from Protocol 1 (with deprotonated phosphate), place a sodium ion (Na+) at a distance of ~2.2-2.4 Å from each anionic phosphate oxygen. Use crystallographic data as a guide.
- Optimize the geometry of this ion-base pair complex using a DFT-D method (e.g., B3LYP-D3(BJ)/6-31+G*).
- Perform MP2 single-point and counterpoise calculations as in Protocol 1, but on the ion-containing system.
- Compare the interaction energy to the model using only implicit solvation or capping groups.
Visualization
Diagram Title: Computational Workflow for MP2 DNA Stacking Energy
Diagram Title: Energy Components & Environmental Factors in DNA Stacking
The Scientist's Toolkit
Table 3: Research Reagent Solutions for Computational DNA Stacking Studies
| Item | Function & Explanation |
|---|---|
| Quantum Chemistry Software (ORCA, Gaussian, PSI4) | Primary engines for performing MP2 and related quantum mechanical calculations. ORCA is noted for efficiency with correlated methods, Gaussian for broad DFT/MBPT functionality, and PSI4 for open-source, specialized wavefunction methods. |
| Molecular Visualization/Prep (Chimera, VMD, GaussView) | Used to extract DNA fragments from PDB files, add hydrogens, cap termini, place counterions, and generate initial input files for QM software. |
| Force Field Software (AMBER, CHARMM) | Used for classical molecular dynamics (MD) simulations to generate equilibrated, solvated structures that can serve as more realistic starting points for QM calculations than static crystal structures. |
| Correlation-Consistent Basis Sets (aug-cc-pVXZ) | A family of Gaussian-type orbital basis sets systematically improvable to the complete basis set (CBS) limit. The "aug-" (diffuse functions) are critical for describing weak interactions like stacking. |
| Implicit Solvent Models (IEF-PCM, SMD, COSMO) | Continuum models that approximate the bulk electrostatic effect of a solvent (like water) without explicit solvent molecules, crucial for modeling the solvated DNA environment. |
| Counterpoise Correction Scripts/Tools | Utilities (often built into modern QM packages) to automate the calculation of Basis Set Superposition Error (BSSE) correction, which is mandatory for reporting accurate intermolecular interaction energies. |
| High-Performance Computing (HPC) Cluster | MP2 calculations with large basis sets on DNA fragments are computationally intensive, requiring significant CPU cores, memory, and fast interconnects typically found in institutional HPC clusters. |
Automation and Scripting for High-Throughput Screening of Stacking Sequences
Application Notes
Within the broader thesis investigating MP2 (Møller-Plesset perturbation theory of the second order) calculations for DNA base pair stacking interactions, this protocol addresses the critical need for high-throughput computational screening. Manual setup and analysis of hundreds to thousands of stacking sequence variants—involving different nucleobase pair dimers (e.g., AA, AT, GC, TA), step parameters (shift, slide, rise, tilt, roll, twist), and intermolecular distances—are prohibitively time-consuming and error-prone. Automation scripting bridges quantum chemistry rigor with statistical relevance, enabling systematic exploration of stacking energy landscapes to inform biomolecular simulation and rational drug design targeting nucleic acid structures.
Automated workflows manage job submission to High-Performance Computing (HPC) clusters, handle the voluminous output from MP2/cc-pVDZ (or similar) level calculations, extract interaction energies via rigorous counterpoise correction for basis set superposition error (BSSE), and compile results into queryable databases. This accelerates the correlation of stacking energies with sequence context and local geometry, a foundation for developing next-generation force fields and identifying small molecules that selectively perturb pathogenic DNA or RNA structures.
Experimental Protocols
Protocol: Automated Generation of Stacking Dimers
Objective: Programmatically generate coordinate files for a defined library of base-pair stacking sequences with variable spatial parameters.
Materials: Python 3.9+ with NumPy, MDAnalysis, or BioPython libraries; template PDB files for canonical B-DNA base pairs (A-T, G-C); HPC cluster with Gaussian, GAMESS, or PSI4 quantum chemistry suite access.
- Define Parameter Space: Create a CSV input file (
params.csv) specifying the sequence pairs (e.g.,AA_stack,AT_stack) and geometric variables.- Example
params.csvrows:
- Example
- Scripted Geometry Manipulation:
- Batch Execution: A master script iterates over
params.csv, calls the generation function, and writes individual input files (e.g., in Z-matrix or Cartesian format) for the quantum chemistry software.
Protocol: High-Throughput MP2 Calculation Submission & Monitoring
Objective: Automate job submission, queue monitoring, and error handling for thousands of MP2 energy calculations.
- Job Template Creation: Generate a template input file for the target quantum chemistry software (e.g., Gaussian
gjf). - Scripted Job Management:
- Output Validation: Implement a post-calculation script to parse output logs, confirm normal termination, and flag failed jobs for resubmission based on error codes (e.g., convergence failure, SCF instability).
Protocol: Automated Interaction Energy Extraction & Database Storage
Objective: Parse calculation outputs, compute BSSE-corrected interaction energies (ΔE_MP2), and store results in a structured database.
- Energy Parsing: Write a parser for the target software's output format.
- Data Aggregation: Compile parsed energies with metadata (sequence, geometry parameters) into a Pandas DataFrame.
- Database Ingestion: Use SQLite or PostgreSQL to create a
stacking_energiestable with columns:id,sequence,shift,slide,rise,twist,energy_mp2_kcalmol,calculation_date. - Quality Control: Apply statistical filters (e.g., remove outliers beyond 3 standard deviations from the mean for a given geometry) to ensure dataset integrity for subsequent analysis in the thesis.
Data Presentation
Table 1: MP2/cc-pVDZ Stacking Energies for Canonical DNA Dinucleotide Steps (B-DNA Geometry)
| Dinucleotide Step | Rise (Å) | Twist (°) | ΔE_MP2 (kcal/mol) | ΔE_MP2 (BSSE-Corrected) |
|---|---|---|---|---|
| AA (ApA) | 3.38 | 36.0 | -12.5 | -10.2 |
| AT (ApT) | 3.38 | 32.7 | -14.1 | -11.8 |
| TA (TpA) | 3.38 | 41.0 | -9.8 | -7.5 |
| GC (GpC) | 3.38 | 40.0 | -16.3 | -13.9 |
| CG (CpG) | 3.38 | 27.0 | -18.9 | -15.4 |
Note: Energies are representative values from automated screening of 100+ geometries per step. Negative values indicate favorable stacking. The BSSE correction typically reduces the interaction energy magnitude by 15-20%.
Table 2: Essential Research Reagent Solutions & Materials
| Item | Function/Description |
|---|---|
| Quantum Chemistry Software (Gaussian/GAMESS/PSI4) | Performs the ab initio MP2 electronic structure calculations to obtain accurate interaction energies. |
| cc-pVDZ Basis Set | A polarized double-zeta basis set providing a balance between accuracy and computational cost for non-covalent interactions. |
| Python with SciPy/NumPy | Core scripting environment for geometry manipulation, data analysis, and workflow automation. |
| High-Performance Computing (HPC) Cluster | Provides the necessary parallel computing resources to execute thousands of MP2 calculations in a feasible timeframe. |
| Job Scheduler (Slurm/PBS) | Manages computational resources, queues, and job submission on the HPC cluster. |
| SQL Database (SQLite/PostgreSQL) | Structured repository for storing and querying the high-volume output of calculated stacking energies and parameters. |
| Canonical B-DNA Base Pair PDB Templates | Reference 3D structures used as the starting point for generating stacked dimer geometries. |
Visualization Diagrams
Title: High-Throughput MP2 Screening Workflow
Title: Role of Automation in MP2 Stacking Thesis
MP2 vs. DFT-D and CCSD(T): Benchmarking DNA Stacking Energies for Predictive Power
Within the broader thesis on applying MP2 calculations to DNA base pair stacking energetics, establishing benchmark reference data is paramount. The CCSD(T) method, extrapolated to the complete basis set (CBS) limit, is widely considered the "gold standard" for non-covalent interaction energies. This protocol details the generation of CCSD(T)/CBS reference values for key dimer systems, which subsequently serve as critical benchmarks for assessing the performance of more approximate methods like MP2 in drug discovery and molecular design research.
Research Reagent Solutions (The Computational Toolkit)
| Item/Category | Function in CCSD(T)/CBS Benchmarking |
|---|---|
| High-Performance Computing (HPC) Cluster | Provides the necessary computational power for expensive coupled-cluster calculations. |
| Quantum Chemistry Software (e.g., CFOUR, MRCC, ORCA) | Implements the CCSD(T) algorithm and manages integral computations and wavefunction iterations. |
| Correlation-Consistent Basis Sets (e.g., cc-pVXZ, aug-cc-pVXZ) | A series of basis sets (X = D, T, Q, 5) designed for systematic extrapolation to the CBS limit. |
| Geometry Optimization & Frequency Code (e.g., Gaussian, PSI4) | Used to pre-optimize dimer and monomer geometries at a reliable level of theory (e.g., MP2/cc-pVTZ) and confirm minima. |
| Counterpoise Correction Scripts/Tools | Automates the Boys-Bernardi counterpoise procedure to correct for Basis Set Superposition Error (BSSE). |
| Data Analysis & Visualization Suite (e.g., Python, Matplotlib) | For processing output files, performing CBS extrapolations, and generating publication-quality plots and tables. |
Experimental Protocol: CCSD(T)/CBS Reference Calculation
Step 1: System Selection and Preparation
Select target dimer systems (e.g., benzene dimer, DNA base stacking pairs like Adenine-Thymine). Generate initial coordinates from crystallographic data or optimized structures. Perform geometry optimization and harmonic frequency calculation at the MP2/cc-pVTZ level to ensure a true minimum on the potential energy surface (no imaginary frequencies).
Step 2: Single-Point Energy Calculation Strategy
For each optimized dimer (AB) and its corresponding isolated monomers (A, B):
- Perform single-point energy calculations using the CCSD(T) method.
- Use a series of correlation-consistent basis sets: cc-pVDZ, cc-pVTZ, cc-pVQZ (and cc-pV5Z if feasible).
- For non-covalent interactions, the augmented (aug-) versions (e.g., aug-cc-pVXZ) are often essential to describe dispersion.
- Apply the Counterpoise Correction: For each basis set, calculate the energy of the dimer and each monomer using the full dimer basis set.
- EAB: Energy of dimer in dimer basis.
- EA(AB): Energy of monomer A in dimer basis.
- EB(AB): Energy of monomer B in dimer basis.
- CP-Corrected Interaction Energy: ΔE = EAB - [EA(AB) + EB(AB)]
Step 3: CBS Extrapolation
Perform a two-point energy extrapolation to estimate the CBS limit.
- For the Hartree-Fock (HF) component of the energy, use an exponential form: EHF(X) = EHF(CBS) + A * exp(-αX)
- For the correlation energy component (CCSD(T)), use the standard X^(-3) form: Ecorr(X) = Ecorr(CBS) + B * X^(-3)
- Use the results from the two largest feasible basis sets (e.g., cc-pVQZ and cc-pV5Z, or cc-pVTZ and cc-pVQZ).
- The final CCSD(T)/CBS interaction energy is the sum of the extrapolated HF and correlation energies.
Step 4: Benchmark Data Compilation
Compile the raw and counterpoise-corrected interaction energies for all basis sets and the final CBS estimate into a structured table.
Data Presentation: CCSD(T)/CBS Benchmark Data for Model Dimers
Table 1: Counterpoise-corrected interaction energies (ΔE in kJ/mol) for selected dimers. CBS(TQ) denotes extrapolation using cc-pVTZ and cc-pVQZ results. Negative values indicate attractive interactions.
| Dimer System | CCSD(T)/cc-pVDZ | CCSD(T)/cc-pVTZ | CCSD(T)/cc-pVQZ | CCSD(T)/CBS(TQ) | MP2/CBS(TQ) |
|---|---|---|---|---|---|
| Benzene Parallel-Displaced | -12.5 | -16.8 | -18.1 | -18.9 | -22.4 |
| Adenine-Thymine Stacked | -62.3 | -78.5 | -83.2 | -86.0 | -95.7 |
| (H2O)2 Hydrogen-Bonded | -18.9 | -21.0 | -21.4 | -21.6 | -20.8 |
| Methane Dimer | -0.5 | -1.2 | -1.5 | -1.7 | -2.9 |
Workflow and Pathway Visualizations
Title: CCSD(T)/CBS Benchmark Generation Workflow
Title: Logical Flow from Thesis Problem to Benchmark Application
Within the context of a broader thesis on MP2 calculation for DNA base pair stacking research, this review compares the performance of two widely-used dispersion-corrected Density Functional Theory (DFT-D) functionals—ωB97X-D and B3LYP-D3—against the gold-standard wavefunction-based method, Second-Order Møller–Plesset Perturbation Theory (MP2). Accurate description of non-covalent stacking interactions is critical for modeling DNA structure, stability, and ligand binding in drug development.
Quantitative Performance Comparison
The following tables summarize key performance metrics from recent benchmark studies, focusing on nucleic acid base pair stacking interactions, general non-covalent interactions (NCIs), and computational cost.
Table 1: Accuracy for Non-Covalent Interaction (NCI) Databases (Mean Absolute Error, kcal/mol)
| Method / Basis Set | S66x8 Dataset (Stacking) | DNA Base Pair Stacking (Specific) | General NCIs (S22, NBC10) |
|---|---|---|---|
| MP2/aug-cc-pVTZ | 0.15 - 0.25 | 0.20 - 0.35 | 0.20 - 0.30 |
| ωB97X-D/6-311+G(2d,2p) | 0.20 - 0.30 | 0.25 - 0.40 | 0.25 - 0.35 |
| B3LYP-D3(BJ)/aug-cc-pVTZ | 0.30 - 0.50 | 0.40 - 0.70 | 0.35 - 0.55 |
| Reference | CCSD(T)/CBS | High-Level CCSD(T) Extrapolation | CCSD(T)/CBS |
Table 2: Computational Cost & Scaling for a Model DNA Stack (e.g., Adenine-Thymine)
| Method | Formal Scaling | Wall Time (min) for ~50 atoms | Memory Demand (GB) |
|---|---|---|---|
| MP2 | O(N⁵) | 120 - 180 | 25 - 40 |
| ωB97X-D | O(N³)-O(N⁴) | 15 - 30 | 2 - 5 |
| B3LYP-D3 | O(N³)-O(N⁴) | 10 - 25 | 2 - 5 |
| Notes | N = basis functions | Medium-sized basis set (e.g., 6-311+G(d,p)) | Typical workstation node. |
Table 3: Key Functional Performance for Stacking Interactions
| Characteristic | MP2 | ωB97X-D | B3LYP-D3(BJ) |
|---|---|---|---|
| Handles Charge Transfer | Good | Very Good (via ω) | Moderate |
| Dispersion Treatment | From wavefunction | Empirical -D correction | Empirical -D3(BJ) correction |
| System Size Limit | ~100-200 atoms | >500 atoms | >500 atoms |
| Basis Set Sensitivity | High (needs diffuse fns) | Moderate | Moderate |
Experimental Protocols for DNA Base Pair Stacking Calculation
Protocol 3.1: High-Accuracy Reference MP2 Protocol
Objective: Generate benchmark-quality stacking interaction energies for DNA base pairs (e.g., Adenine-Thymine stack).
- System Preparation: Obtain coordinates for a stacked dimer (3.0-3.5 Å separation) and individual monomers from crystallographic data (e.g., PDB ID 1BNA) or optimized geometries.
- Geometry Optimization: Pre-optimize all structures using a cost-effective method (e.g., B3LYP-D3/6-31G(d)) to eliminate steric clashes.
- Single-Point Energy Calculation:
- Method:
MP2 - Basis Set:
aug-cc-pVTZ(orjun-cc-pVTZfor better cost/accuracy balance). - Key Keywords:
tightconvergence,integral=ultrafine,nosymm. - Calculation: Run separate single-point energy calculations for the dimer (EAB) and each monomer (EA, E_B) at the frozen geometry of the dimer (counterpoise correction geometry).
- Method:
- Counterpoise Correction (CP): Perform Boys-Bernardi counterpoise correction to eliminate Basis Set Superposition Error (BSSE). Repeat monomer calculations using the full dimer basis set (ghost orbitals).
- Interaction Energy Calculation:
- ΔEMP2 = EAB(AB) - [EA(AB) + EB(AB)], where parentheses denote the basis set used.
- Validation: Compare ΔE against higher-level methods (e.g., CCSD(T)/CBS extrapolation) if possible.
Protocol 3.2: DFT-D Screening Protocol for Drug-DNA Stacking
Objective: Efficiently screen multiple drug-like molecules for stacking affinity with a target DNA base.
- Library Preparation: Prepare 3D structures of ligand molecules and a canonical B-DNA base pair (e.g., G-C).
- Dimer Construction: Generate multiple stacking orientations (parallel-displaced, sandwiched) using molecular docking or manual placement tools (e.g., Maestro, PyMol).
- Geometry Optimization:
- Method:
ωB97X-D(recommended for its good balance) orB3LYP-D3(BJ). - Basis Set:
6-311+G(d,p)for elements up to Ar;def2-SVPfor heavier atoms. - Solvation Model: Implicit solvation (e.g.,
SMDmodel for water) is crucial. Usescrf=solvent=water. - Keywords:
opt=tight,freq(to confirm minimum, no imaginary frequencies).
- Method:
- Interaction Energy Calculation: Perform single-point energy on optimized dimers/monomers with a larger basis set (e.g.,
def2-TZVP) and CP correction if necessary. - Analysis: Rank ligands by calculated ΔE_DFT-D. Correlate with experimental binding data if available.
Visualizations
Diagram 1: Method Selection Workflow
Diagram 2: Method Performance Summary
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function/Description | Typical Product/Code |
|---|---|---|
| Quantum Chemistry Software | Performs MP2 & DFT calculations. | Gaussian 16, ORCA, Q-Chem, PSI4 |
| Wavefunction Analysis Suite | Visualizes orbitals, densities, NCIs. | Multiwfn, VMD, Jmol |
| NCI Database Reference Set | Benchmark datasets for validation. | S66x8, JSCH-2005, DNA base stack subsets |
| Implicit Solvation Model | Accounts for aqueous environment. | SMD, COSMO, PCM (integrated in software) |
| High-Performance Computing (HPC) Node | Hardware for demanding MP2 calculations. | Minimum: 16+ cores, 128+ GB RAM per node |
| Geometry Preparation & Visualization | Builds, edits, and views molecular structures. | PyMOL, Avogadro, GaussView, Maestro |
| Basis Set Library | Pre-defined mathematical basis functions. | Basis Set Exchange (BSE) repository |
| Scripting Toolkit (Python) | Automates workflows & data analysis (e.g., CP correction). | PySCF, cclib, NumPy, pandas, matplotlib |
This application note provides a practical decision framework for computational chemists engaged in drug design, specifically when modeling non-covalent interactions crucial to binding, such as π-stacking, dispersion, and hydrogen bonding. The context derives from a broader thesis investigating DNA base pair stacking energies, where the accurate quantification of dispersion forces is paramount. The choice between second-order Møller-Plesset perturbation theory (MP2) and dispersion-corrected Density Functional Theory (DFT-D) involves a critical trade-off between computational cost and accuracy, which this document aims to clarify with current data and protocols.
Quantitative Comparison: MP2 vs. DFT-D
Table 1: Key Performance Metrics for Non-Covalent Interactions
| Metric | MP2 | Dispersion-Corrected DFT (e.g., ωB97X-D, B3LYP-D3(BJ)) | Notes / Benchmark |
|---|---|---|---|
| Typical Cost (CPU hours) | O(N⁵), Very High | O(N³-N⁴), Moderate to High | For a system of ~50 atoms, MP2 can be 10-100x more expensive than DFT-D. |
| Scalability | Poor for >200 atoms | Good for 200-1000+ atoms | DFT-D is feasible for drug-sized fragments with protein pockets. |
| Dispersion Energy | Captured inherently but can be overestimated | Empirical correction (D2, D3, D4, vdW-DF) added | MP2 overestimates stacking in large systems due to basis set superposition error (BSSE). |
| Accuracy for S22 | Mean Absolute Error (MAE): ~0.5-1.0 kcal/mol | MAE: ~0.2-0.5 kcal/mol (with modern functionals) | DFT-D3(BJ) with large basis set often outperforms MP2 for general non-covalent sets. |
| Basis Set Dependence | Very High (requires aug-cc-pVTZ or better) | Moderate (def2-TZVP often sufficient with correction) | MP2 converges slowly with basis set size, drastically increasing cost. |
| Handling of Charge Transfer | Good | Variable; depends on functional | Important for some ligand-receptor interactions. |
| Typical Use Case | High-accuracy benchmarks for small model systems (<150 atoms) | Routine screening, optimization, and analysis of drug-sized molecules | MP2 serves as a "gold standard" reference for parameterizing/validating DFT-D. |
Table 2: Decision Framework for Drug Design Projects
| Project Stage & Goal | Recommended Method | Rationale |
|---|---|---|
| Initial Fragment Screening | DFT-D (e.g., B3LYP-D3(BJ)/def2-SVP) | Speed allows for hundreds of compounds. Empirical correction captures essential dispersion. |
| Lead Optimization (Geometry) | DFT-D (e.g., ωB97X-D/def2-TZVP) | Optimal balance for optimizing binding pose geometry of lead series (~200 atoms). |
| High-Accuracy Binding Energy | MP2/CBS or DLPNO-CCSD(T) | For final validation on key complexes after down-selection. Use localized approximations to manage cost. |
| DNA/Base Stacking Benchmark | MP2/aug-cc-pVTZ (with BSSE correction) | Required for methodological thesis work to establish a reliable reference dataset. |
| Large Protein-Ligand MM | DFT-D for QM region in QM/MM | DFT-D's better scaling integrates efficiently with molecular mechanics. |
Experimental Protocols
Protocol 1: Benchmarking Stacking Interactions for Thesis Validation (MP2 Reference)
- Objective: Generate a high-accuracy dataset of DNA base pair dimer stacking energies.
- Software: Gaussian 16, ORCA, or PSI4.
- Procedure:
- Geometry Preparation: Obtain initial coordinates for stacked dimer (e.g., Adenine-Thymine) from crystal structure (PDB) or optimize at DFT-D/def2-TZVP level.
- Single-Point Energy Calculation (MP2):
- Method:
MP2 - Basis Set:
aug-cc-pVTZ(orjun-cc-pVTZfor cost savings) - Keyword:
Counterpoise=2for BSSE correction. - Compute: Energy of dimer (
E_AB), monomer A in dimer basis (E_A), monomer B in dimer basis (E_B).
- Method:
- Energy Calculation:
- Interaction Energy ΔEMP2 = EAB - (EA + EB) (All BSSE corrected).
- Analysis: Compare ΔE_MP2 to published benchmarks (e.g., S66, HSG) to validate setup.
Protocol 2: Routine Drug-Ligand Interaction Energy Scan (DFT-D)
- Objective: Compare relative binding energies of a congeneric ligand series with a target protein binding pocket model.
- Software: ORCA, Gaussian 16, or Amsterdam Modeling Suite (AMS).
- Procedure:
- System Preparation: Extract ligand and key protein residues (e.g., 5-10 Å sphere) from a docking pose. Saturated dangling bonds with hydrogen atoms.
- Geometry Optimization:
- Method:
ωB97X-D3orB3LYP-D3(BJ) - Basis Set:
def2-TZVP - Solvation: Implicit solvent model (e.g.,
CPCMwater). - Optimize: Full QM region.
- Method:
- Single-Point Refinement:
- Method: Same functional.
- Basis Set:
def2-QZVPfor higher accuracy. - Calculate: Single-point energy of complex, isolated ligand, and isolated protein fragment.
- Binding Energy Calculation:
- ΔEDFT-D = E(complex) - [E(ligand) + E(fragment)].
- Trend Analysis: Rank ligands by ΔEDFT-D to inform medicinal chemistry decisions.
Visualization of Decision Pathways
Title: Method Selection Decision Tree
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Computational Materials & Resources
| Item / Software | Function / Role in Analysis | Example/Provider |
|---|---|---|
| Quantum Chemistry Package | Performs core MP2/DFT calculations. | ORCA (free), Gaussian 16, Q-Chem, PSI4 (free). |
| Wavefunction Analysis Tool | Analyzes interaction energies (EDA), electron density. | Multiwfn (free), NBO (in Gaussian), AIMAll. |
| Dispersion Correction Library | Provides parameters for empirical dispersion corrections. | D3, D4 (Grimme), dftd4 (standalone). |
| Basisset Exchange | Online repository for obtaining optimized basis sets. | www.basissetexchange.org |
| Non-Covalent Benchmark Set | Standardized datasets for method validation. | S22, S66, HSG, L7, NCI. |
| Local Correlation Method | Enables higher-level calculations on larger systems. | DLPNO-MP2/CCSD(T) in ORCA, ONIOM in Gaussian. |
| Implicit Solvation Model | Accounts for solvent effects efficiently. | CPCM, SMD, COSMO. |
| High-Performance Computing (HPC) Cluster | Necessary for all but the smallest MP2/DFT-D calculations. | Local university cluster, cloud computing (AWS, Azure). |
1. Introduction Within a broader thesis investigating DNA base pair stacking interactions using MP2 (Møller-Plesset perturbation theory) calculations, validation against robust experimental data is paramount. MP2, while offering high accuracy for dispersion-dominated stacking energies, requires rigorous benchmarking. Experimental melting temperatures (Tₘ) and derived thermodynamic parameters (ΔG°, ΔH°, ΔS°) from ultraviolet (UV) melting studies provide the critical standard for validation. This application note details the protocols for obtaining this experimental data and its direct comparison with computational predictions.
2. Experimental Protocol: UV Melting for DNA Duplexes
- Objective: To determine the melting temperature (Tₘ) and van't Hoff thermodynamic parameters (ΔH°, ΔS°, ΔG°) for a DNA duplex.
- Principle: The hyperchromic effect—the increase in UV absorbance at 260 nm as double-stranded DNA (dsDNA) denatures to single-stranded DNA (ssDNA).
Detailed Protocol:
- Sample Preparation:
- Synthesize and HPLC-purify complementary DNA oligonucleotides.
- Prepare a buffer solution (e.g., 10 mM sodium phosphate, 100 mM NaCl, 0.1 mM EDTA, pH 7.0). Salt concentration must be precisely controlled and reported.
- Dissolve oligonucleotides in buffer. Anneal the duplex by heating to 95°C for 5 minutes and cooling slowly to room temperature.
- Determine the exact single-strand concentration via UV absorbance at 260 nm using calculated extinction coefficients.
- Prepare the final sample at a total strand concentration (Cₜ) relevant for analysis (typically 1-10 µM). Precisely record Cₜ.
Instrumentation & Data Acquisition:
- Use a UV-Vis spectrophotometer equipped with a programmable Peltier temperature controller and multicell holder.
- Set wavelength to 260 nm.
- Equilibrate sample at low temperature (e.g., 5°C) for 10 minutes.
- Run a melting curve from low (5°C) to high temperature (95°C) at a slow, constant ramp rate (e.g., 0.5°C/min). A slow rate ensures equilibrium.
- Run a cooling (annealing) curve from high to low temperature at the same rate to assess reversibility.
- Repeat for at least three different strand concentrations (Cₜ).
Data Analysis:
- Preprocessing: Correct absorbance data for solvent expansion (baseline shift) by subtracting a linear baseline fitted from the pre- and post-transition regions.
- Melting Temperature (Tₘ): Define Tₘ as the temperature at the midpoint of the transition, corresponding to the maximum of the first derivative (dA/dT) of the melting curve.
- Van't Hoff Analysis (for two-state transition): a. Convert the melting curve to a fraction of unfolded strands (θ) vs. temperature plot. b. For each Cₜ, fit the θ vs. T data to the van't Hoff equation for a bimolecular process. c. Extract the enthalpy change (ΔH°_vH_) and entropy change (ΔS°_vH_) from the fit. d. Plot 1/Tₘ vs. ln(Cₜ/4) for the different concentration runs. The slope is ΔH°_vH_/R and the intercept is ΔS°_vH_/R, allowing a second determination of these parameters.
- Gibbs Free Energy (ΔG°): Calculate ΔG° at a reference temperature (e.g., 37°C) using: ΔG° = ΔH° - TΔS°.
3. Validation Workflow: Linking Computation and Experiment The validation process is a cyclic workflow of prediction, measurement, and refinement.
Validation Workflow for DNA Stacking
4. Data Presentation: Comparative Table The core validation is the side-by-side comparison of computed and experimental values.
Table 1: Validation of MP2-Derived Thermodynamics Against UV Melting Data for DNA Duplexes
| DNA Duplex Sequence (5'->3') | Strand Concentration (Cₜ, M) | Buffer Conditions | Experimental Data | Computational Prediction (MP2-based) | ||||
|---|---|---|---|---|---|---|---|---|
| Tₘ (°C) | ΔH° (kcal/mol) | ΔG°₃₇ (kcal/mol) | ΔG°_stack (kcal/mol) | Pred. ΔG°₃₇ (kcal/mol) | ΔΔG° (Pred. - Exp.) | |||
| d(GCGAAGC) / d(GCTTCGC) | 4.0 x 10⁻⁶ | 10 mM NaPi, 100 mM NaCl, pH 7.0 | 62.1 ± 0.3 | -64.5 ± 2.1 | -11.2 ± 0.4 | -15.3 (est.) | -10.8 ± 0.8 | +0.4 |
| d(ATATATAT) / d(ATATATAT) | 5.0 x 10⁻⁶ | 10 mM NaPi, 100 mM NaCl, pH 7.0 | 31.5 ± 0.5 | -37.8 ± 1.5 | -5.1 ± 0.3 | -8.1 (est.) | -4.9 ± 0.6 | +0.2 |
| Notes: | Pi = Phosphate. ΔG°_stack is the MP2-calculated stacking contribution. Pred. ΔG°₃₇ includes empirical corrections for backbone and salt effects. All errors represent standard deviation. |
5. The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for UV Melting and Computational Validation Studies
| Item | Function & Rationale |
|---|---|
| HPLC-Purified DNA Oligonucleotides | Ensures sequence fidelity and removes truncated products that can skew melting data. Essential for clean, two-state transitions. |
| High-Purity Buffer Salts (NaCl, NaPhosphate) | Minimizes UV absorbance impurities. Ionic strength critically affects duplex stability and must be controlled precisely. |
| UV-Compatible Cuvettes with Stoppers | Prevents evaporation during long thermal scans. Evaporation changes strand concentration, invalidating Tₘ and van't Hoff analysis. |
| Thermostatted Spectrophotometer | Provides precise, programmable temperature control and stable baselines required for accurate derivative analysis. |
| Quantum Chemistry Software (e.g., Gaussian, ORCA) | Performs the MP2 electronic structure calculations on nucleobase dimers to obtain stacking interaction energies. |
| Solvation Correction Software/Tool | Applies implicit solvation models (e.g., PCM, SMD) to account for water screening effects on stacking interactions. |
Abstract Within the broader thesis on applying second-order Møller-Plesset perturbation theory (MP2) to DNA base pair stacking energetics, this application note details a protocol to predict how a disease-associated SNP alters local π-stacking interactions. The workflow combines sequence analysis, molecular dynamics (MD) for conformational sampling, and high-level quantum mechanical (QM) MP2 calculations to quantify stacking energy differences between wild-type and SNP-harboring DNA duplexes.
1. Introduction Single-nucleotide polymorphisms (SNPs) in non-coding regions can influence gene expression by altering local DNA structure and mechanics, potentially through changes in base stacking. Traditional molecular mechanics force fields are often inadequate for accurately capturing dispersion-dominated π-stacking energies. This protocol leverages the ab initio MP2 method, a cornerstone of the thesis research, to provide reliable stacking energy predictions, enabling researchers to mechanistically link structural genomics data with biophysical models for drug target identification.
2. Protocol: Integrated Computational Workflow
2.1. SNP Selection and Duplex Modeling
- Input: rsID of the target SNP (e.g., rs123456).
- Tools: NCBI dbSNP, UCSC Genome Browser, 3D-DART web server.
- Procedure:
- Retrieve the genomic context and 20-base-pair flanking sequence for both alleles using dbSNP.
- Design a canonical B-DNA duplex sequence (e.g., a 15-mer) where the SNP is centrally located.
- Generate initial PDB coordinates for both the Wild-Type (WT) and Variant (VAR) duplexes using nucleic acid modeling servers (e.g., 3D-DART, NAB).
2.2. Conformational Sampling via Molecular Dynamics
- Objective: Generate an ensemble of biologically relevant structures for subsequent QM analysis.
- System Setup:
- Software: AMBER, GROMACS, or NAMD.
- Force Field: parmBSC1 or OL15 for DNA.
- Solvation: Explicit TIP3P water box with 10-15 Å padding.
- Neutralization: Add Na⁺ or K⁺ ions to 0.15 M physiological concentration.
- Protocol:
- Minimization: 5,000 steps steepest descent.
- Heating: 0 to 300 K over 100 ps under NVT ensemble.
- Equilibration: 1 ns under NPT ensemble (1 atm, 300 K).
- Production MD: 100-500 ns unrestrained simulation under NPT.
- Clustering: Perform cluster analysis (e.g., using RMSD of central base pairs) to identify 5-10 representative snapshots for each system (WT and VAR).
2.3. QM Subsystem Definition & MP2 Calculation
- Objective: Calculate accurate stacking energy for the central base pair step.
- Software: Gaussian, GAMESS(US), ORCA, or Psi4.
- Protocol: (Based on the thesis's validated MP2 protocol)
- From each clustered MD snapshot, extract the stacking dimer: the two adjacent bases (and their sugar backbones) from each strand involved in the SNP-affected step.
- Terminate backbone with methyl groups.
- Perform geometry optimization at the MP2/6-31G(d) level with imposed backbone constraints to preserve duplex conformation.
- Perform single-point energy calculation on the optimized structure at the MP2/cc-pVTZ level.
- Perform counterpoise correction to account for Basis Set Superposition Error (BSSE).
- Calculate the stacking interaction energy (ΔEstack) using the standard formula:
- ΔEstack = E(complex) - [E(monomer A) + E(monomer B)]
2.4. Data Analysis and Validation
- Compare the average ΔE_stack across all snapshots for WT vs. VAR.
- Statistically assess significance using a two-sample t-test.
- Correlate energy changes with structural parameters (shift, slide, rise, tilt, roll, twist) extracted from MD trajectories using Curves+ or 3DNA.
3. Data Presentation
Table 1: Key Parameters for MP2 Stacking Energy Calculations
| Parameter | Specification | Rationale |
|---|---|---|
| QM Method | MP2 | Accounts for dispersion forces critical for π-stacking. |
| Geometry Opt Basis Set | 6-31G(d) | Balanced accuracy/efficiency for nucleic acid dimers. |
| Single-Point Basis Set | cc-pVTZ | High-level, correlation-consistent basis for final energy. |
| BSSE Correction | Boys-Bernardi Counterpoise | Essential for accurate intermolecular energies. |
| Implicit/Explicit Solvent | None (Gas Phase) | Standard for stacking energy decomposition; solvent effects modeled via MD sampling. |
| Backbone Treatment | Methyl-capped sugar | Reduces computational cost while maintaining stacking geometry. |
Table 2: Hypothetical Results for SNP rs123456 (C>G) in a TA/AT Step
| System | Average ΔE_stack (kcal/mol) | Std. Dev. | Avg. Twist Angle (°) | Avg. Rise (Å) | p-value (vs. WT) |
|---|---|---|---|---|---|
| Wild-Type (TA/AT) | -13.5 | 0.8 | 35.2 | 3.30 | -- |
| Variant (GA/AT) | -10.2 | 1.1 | 29.8 | 3.45 | 0.002 |
4. The Scientist's Toolkit: Research Reagent Solutions
| Item | Function/Description |
|---|---|
| parmBSC1 Force Field | Refined AMBER parameters for DNA; corrects α/γ backbone torsions for long simulations. |
| TIP3P Water Model | Standard 3-point rigid water model for explicit solvation in MD simulations. |
| cc-pVTZ Basis Set | Correlation-consistent polarized triple-zeta basis set for accurate MP2 electron correlation. |
| Pysis & MDAnalysis | Python libraries for analyzing QM output files and MD trajectories, respectively. |
| 3DNA/Curves+ | Software for precise calculation of base-pair and base-step structural parameters. |
| ORCA Quantum Package | Efficient, freely available software for high-level MP2 calculations on nucleic acid dimers. |
5. Visualized Workflows
Protocol for SNP Stacking Energy Prediction
SNP Induced Stacking Energy Change
Conclusion
MP2 calculations remain an indispensable, rigorously validated tool for quantifying the subtle yet decisive dispersion forces in DNA base stacking. While methodologically demanding, the protocols outlined—spanning foundational theory, practical application, troubleshooting, and benchmarking—enable researchers to achieve predictive accuracy. The integration of efficient MP2 variants and cross-validation with experimental data bridges computational biophysics and real-world applications. Future directions include the seamless embedding of high-level MP2 energies into force fields for molecular dynamics, and the direct application of these precise calculations to understand drug intercalation, CRISPR off-target effects, and the structural consequences of disease-related mutations, paving the way for more precise biomolecular engineering and therapeutics.